Abstract
Eukaryotic DNA mismatch repair (MMR) involves both exonuclease 1 (Exo1)-dependent and Exo1-independent pathways. We found that the unstructured C-terminal domain of Saccharomyces cerevisiae Exo1 contains two MutS homolog 2 (Msh2)-interacting peptide (SHIP) boxes downstream from the MutL homolog 1 (Mlh1)-interacting peptide (MIP) box. These three sites were redundant in Exo1-dependent MMR in vivo and could be replaced by a fusion protein between an N-terminal fragment of Exo1 and Msh6. The SHIP-Msh2 interactions were eliminated by the msh2M470I mutation, and wild-type but not mutant SHIP peptides eliminated Exo1-dependent MMR in vitro. We identified two S. cerevisiae SHIP-box-containing proteins and three candidate human SHIP-box-containing proteins. One of these, Fun30, had a small role in Exo1-dependent MMR in vivo. The Remodeling of the Structure of Chromatin (Rsc) complex also functioned in both Exo1-dependent and Exo1-independent MMR in vivo. Our results identified two modes of Exo1 recruitment and a peptide module that mediates interactions between Msh2 and other proteins, and they support a model in which Exo1 functions in MMR by being tethered to the Msh2–Msh6 complex.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Change history
04 November 2019
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Goellner, E. M., Putnam, C. D. & Kolodner, R. D. Exonuclease-1-dependent and independent mismatch repair. DNA Repair 32, 24–32 (2015).
Fishel, R. Mismatch repair. J. Biol. Chem. 290, 26395–26403 (2015).
Li, Z., Pearlman, A. H. & Hsieh, P. DNA mismatch repair and the DNA damage response. DNA Repair 38, 94–101 (2016).
Kolodner, R. D. & Marsischky, G. T. Eukaryotic DNA mismatch repair. Curr. Opin. Genet. Dev. 9, 89–96 (1999).
Li, G. M. Mechanisms and functions of DNA mismatch repair. Cell Res. 18, 85–98 (2008).
Spies, M. & Fishel, R. Mismatch repair during homologous and homeologous recombination. Cold Spring Harb. Perspect. Biol. 7, a022657 (2015).
Lynch, H. T., Snyder, C. L., Shaw, T. G., Heinen, C. D. & Hitchins, M. P. Milestones of Lynch syndrome: 1895–2015. Nat. Rev. Cancer 15, 181–194 (2015).
Durno, C. A. et al. Phenotypic and genotypic characterization of biallelic mismatch repair deficiency (BMMR-D) syndrome. Eur. J. Cancer 51, 977–983 (2015).
Waterfall, J. J. & Meltzer, P. S. Avalanching mutations in biallelic mismatch repair deficiency syndrome. Nat. Genet. 47, 194–196 (2015).
Kane, M. F. et al. Methylation of the hMLH1 promoter correlates with lack of expression of hMLH1 in sporadic colon tumors and mismatch-repair-defective human tumor cell lines. Cancer Res. 57, 808–811 (1997).
Orans, J. et al. Structures of human exonuclease 1 DNA complexes suggest a unified mechanism for nuclease family. Cell 145, 212–223 (2011).
Tishkoff, D. X. et al. Identification and characterization of Saccharomyces cerevisiae EXO1, a gene encoding an exonuclease that interacts with MSH2. Proc. Natl Acad. Sci. USA 94, 7487–7492 (1997).
Genschel, J. & Modrich, P. Mechanism of 5′-directed excision in human mismatch repair. Mol. Cell 12, 1077–1086 (2003).
Bowen, N. et al. Reconstitution of long- and short-patch mismatch repair reactions using Saccharomyces cerevisiae proteins. Proc. Natl Acad. Sci. USA 110, 18472–18477 (2013).
Bowen, N. & Kolodner, R. D. Reconstitution of Saccharomyces cerevisiae DNA polymerase ε–dependent mismatch repair with purified proteins. Proc. Natl Acad. Sci. USA 114, 3607–3612 (2017).
Zhang, Y. et al. Reconstitution of 5′-directed human mismatch repair in a purified system. Cell 122, 693–705 (2005).
Constantin, N., Dzantiev, L., Kadyrov, F. A. & Modrich, P. Human mismatch repair: reconstitution of a nick-directed bidirectional reaction. J. Biol. Chem. 280, 39752–39761 (2005).
Smith, C. E. et al. Activation of Saccharomyces cerevisiae Mlh1–Pms1 endonuclease in a reconstituted mismatch repair system. J. Biol. Chem. 290, 21580–21590 (2015).
Kadyrov, F. A., Dzantiev, L., Constantin, N. & Modrich, P. Endonucleolytic function of MutLα in human mismatch repair. Cell 126, 297–308 (2006).
Amin, N. S., Nguyen, M. N., Oh, S. & Kolodner, R. D. Exo1-dependent mutator mutations: model system for studying functional interactions in mismatch repair. Mol. Cell. Biol. 21, 5142–5155 (2001).
Wei, K. et al. Inactivation of exonuclease 1 in mice results in DNA mismatch repair defects, increased cancer susceptibility, and male and female sterility. Genes Dev. 17, 603–614 (2003).
Shell, S. S., Putnam, C. D. & Kolodner, R. D. The N terminus of Saccharomyces cerevisiae Msh6 is an unstructured tether to PCNA. Mol. Cell 26, 565–578 (2007).
Smith, C. E. et al. Dominant mutations in S. cerevisiae PMS1 identify the Mlh1–Pms1 endonuclease active site and an exonuclease-1-independent mismatch repair pathway. PLoS Genet. 9, e1003869 (2013).
Goellner, E. M. et al. PCNA and Msh2–Msh6 activate an Mlh1–Pms1 endonuclease pathway required for Exo1-independent mismatch repair. Mol. Cell 55, 291–304 (2014).
Hombauer, H., Campbell, C. S., Smith, C. E., Desai, A. & Kolodner, R. D. Visualization of eukaryotic DNA mismatch repair reveals distinct recognition and repair intermediates. Cell 147, 1040–1053 (2011).
Kadyrov, F. A. et al. A possible mechanism for exonuclease-1-independent eukaryotic mismatch repair. Proc. Natl Acad. Sci. USA 106, 8495–8500 (2009).
Schmutte, C., Sadoff, M. M., Shim, K. S., Acharya, S. & Fishel, R. The interaction of DNA mismatch repair proteins with human exonuclease 1. J. Biol. Chem. 276, 33011–33018 (2001).
Dherin, C. et al. Characterization of a highly conserved binding site of Mlh1 required for exonuclease-1-dependent mismatch repair. Mol. Cell. Biol. 29, 907–918 (2009).
Gueneau, E. et al. Structure of the MutLα C-terminal domain reveals how Mlh1 contributes to Pms1 endonuclease site. Nat. Struct. Mol. Biol. 20, 461–468 (2013).
Tran, P. T. et al. A mutation in EXO1 defines separable roles in DNA mismatch repair and post-replication repair. DNA Repair 6, 1572–1583 (2007).
Gellon, L., Werner, M. & Boiteux, S. Ntg2p, a Saccharomyces cerevisiae DNA N-glycosylase and apurinic or apyrimidinic lyase involved in base-excision repair of oxidative DNA damage, interacts with the DNA mismatch repair protein Mlh1p. Identification of a Mlh1p-binding motif. J. Biol. Chem. 277, 29963–29972 (2002).
Umar, A. et al. Requirement for PCNA in DNA mismatch repair at a step preceding DNA resynthesis. Cell 87, 65–73 (1996).
Warren, J. J. et al. Structure of the human MutSα DNA lesion recognition complex. Mol. Cell 26, 579–592 (2007).
Stormo, G. D., Schneider, T. D., Gold, L. & Ehrenfeucht, A. Use of the ‘Perceptron’ algorithm to distinguish translational initiation sites in E. coli. Nucleic Acids Res 10, 2997–3011 (1982).
Huh, W. K. et al. Global analysis of protein localization in budding yeast. Nature 425, 686–691 (2003).
Kumar, A. et al. Subcellular localization of the yeast proteome. Genes Dev. 16, 707–719 (2002).
Natter, K. et al. The spatial organization of lipid synthesis in the yeast Saccharomyces cerevisiae derived from large-scale green fluorescent protein tagging and high-resolution microscopy. Mol. Cell. Proteom. 4, 662–672 (2005).
Gauci, S., Veenhoff, L. M., Heck, A. J. & Krijgsveld, J. Orthogonal separation techniques for the characterization of the yeast nuclear proteome. J. Proteome Res. 8, 3451–3463 (2009).
Campbell, C. S. et al. Mlh2 is an accessory factor for DNA mismatch repair in Saccharomyces cerevisiae. PLoS Genet. 10, e1004327 (2014).
Dosztányi, Z., Csizmok, V., Tompa, P. & Simon, I. IUPred: web server for the prediction of intrinsically unstructured regions of proteins based on estimated energy content. Bioinformatics 21, 3433–3434 (2005).
Awad, S., Ryan, D., Prochasson, P., Owen-Hughes, T. & Hassan, A. H. The Snf2 homolog Fun30 acts as a homodimeric ATP-dependent chromatin-remodeling enzyme. J. Biol. Chem. 285, 9477–9484 (2010).
Chen, X. et al. The Fun30 nucleosome remodeler promotes resection of DNA double-strand break ends. Nature 489, 576–580 (2012).
Costelloe, T. et al. The yeast Fun30 and human SMARCAD1 chromatin remodelers promote DNA end resection. Nature 489, 581–584 (2012).
Araki, H. et al. Cloning DPB3, the gene encoding the third subunit of DNA polymerase II of Saccharomyces cerevisiae. Nucleic Acids Res. 19, 4867–4872 (1991).
Datta, A., Adjiri, A., New, L., Crouse, G. F. & Jinks Robertson, S. Mitotic cross-overs between diverged sequences are regulated by mismatch repair proteins in Saccharomyces cerevisiae. Mol. Cell. Biol. 16, 1085–1093 (1996).
Myung, K., Datta, A., Chen, C. & Kolodner, R. D. SGS1, the Saccharomyces cerevisiae homolog of BLM and WRN, suppresses genome instability and homeologous recombination. Nat. Genet. 27, 113–116 (2001).
Bermudez, V. P., Farina, A., Tappin, I. & Hurwitz, J. Influence of the human cohesion establishment factor Ctf4, AND-1, on DNA replication. J. Biol. Chem. 285, 9493–9505 (2010).
Gambus, A. et al. A key role for Ctf4 in coupling the MCM2–7 helicase to DNA polymerase-α within the eukaryotic replisome. EMBO J. 28, 2992–3004 (2009).
Im, J. S. et al. Assembly of the Cdc45–Mcm2–7–GINS complex in human cells requires the Ctf4 (AND-1), RECQL4 and MCM10 proteins. Proc. Natl. Acad. Sci. USA 106, 15628–15632 (2009).
Zhu, W. et al. Mcm10 and AND-1/CTF4 recruit DNA polymerase-α to chromatin for initiation of DNA replication. Genes Dev. 21, 2288–2299 (2007).
Chen, Z. et al. Proteomic analysis reveals a novel mutator S (MutS) partner involved in mismatch repair pathway. Mol. Cell. Proteom. 15, 1299–1308 (2016).
Traver, S. et al. MCM9 is required for mammalian DNA mismatch repair. Mol. Cell 59, 831–839 (2015).
Jeon, Y. et al. Dynamic control of strand excision during human DNA mismatch repair. Proc. Natl Acad. Sci. USA 113, 3281–3286 (2016).
Myler, L. R. et al. Single-molecule imaging reveals the mechanism of Exo1 regulation by single-stranded DNA-binding proteins. Proc. Natl Acad. Sci. USA 113, E1170–E1179 (2016).
Li, G. M. & Modrich, P. Restoration of mismatch repair to nuclear extracts of H6 colorectal tumor cells by a heterodimer of human MutL homologs. Proc. Natl Acad. Sci. USA 92, 1950–1954 (1995).
Geng, H. et al. In vitro studies of DNA mismatch repair proteins. Anal. Biochem. 413, 179–184 (2011).
Verreault, A. De novo nucleosome assembly: new pieces in an old puzzle. Genes Dev. 14, 1430–1438 (2000).
Hombauer, H., Srivatsan, A., Putnam, C. D. & Kolodner, R. D. Mismatch repair, but not heteroduplex rejection, is temporally coupled to DNA replication. Science 334, 1713–1716 (2011).
Javaid, S. et al. Nucleosome remodeling by hMSH2–hMSH6. Mol. Cell 36, 1086–1094 (2009).
Obmolova, G., Ban, C., Hsieh, P. & Yang, W. Crystal structures of mismatch repair protein MutS and its complex with a substrate DNA. Nature 407, 703–710 (2000).
Sikorski, R. S. & Hieter, P. A system of shuttle vectors and yeast host strains designed for efficient manipulation of DNA in Saccharomyces cerevisiae. Genetics 122, 19–27 (1989).
Yan, D. & Jin, Y. Regulation of DLK-1 kinase activity by calcium-mediated dissociation from an inhibitory isoform. Neuron 76, 534–548 (2012).
Hargreaves, V. V., Shell, S. S., Mazur, D. J., Hess, M. T. & Kolodner, R. D. Interaction between the Msh2 and Msh6 nucleotide-binding sites in the Saccharomyces cerevisiae Msh2–Msh6 complex. J. Biol. Chem. 285, 9301–9310 (2010).
Antony, E. & Hingorani, M. M. Mismatch-recognition-coupled stabilization of Msh2–Msh6 in an ATP-bound state at the initiation of DNA repair. Biochemistry 42, 7682–7693 (2003).
Fien, K. & Stillman, B. Identification of replication factor C from Saccharomyces cerevisiae: a component of the leading-strand DNA replication complex. Mol. Cell. Biol. 12, 155–163 (1992).
Fortune, J. M., Stith, C. M., Kissling, G. E., Burgers, P. M. & Kunkel, T. A. RPA and PCNA suppress formation of large deletion errors by yeast DNA polymerase-δ. Nucleic Acids Res 34, 4335–4341 (2006).
Gomes, X. V., Gary, S. L. & Burgers, P. M. Overproduction in Escherichia coli and characterization of yeast replication factor C lacking the ligase homology domain. J. Biol. Chem. 275, 14541–14549 (2000).
Nakagawa, T., Flores-Rozas, H. & Kolodner, R. D. The MER3 helicase involved in meiotic crossing over is stimulated by single-stranded DNA-binding proteins and unwinds DNA in the 3′ to 5′ direction. J. Biol. Chem. 276, 31487–31493 (2001).
Jenuth, J. P. The NCBI. Publicly available tools and resources on the Web. Methods Mol. Biol. 132, 301–312 (2000).
Cock, P. J. et al. Biopython: freely available Python tools for computational molecular biology and bioinformatics. Bioinformatics 25, 1422–1423 (2009).
Katoh, K. & Standley, D. M. MAFFT: iterative refinement and additional methods. Methods Mol. Biol. 1079, 131–146 (2014).
Scannell, D. R., Butler, G. & Wolfe, K. H. Yeast genome evolution—the origin of the species. Yeast 24, 929–942 (2007).
Retief, J. D. Phylogenetic analysis using PHYLIP. Methods Mol. Biol. 132, 243–258 (2000).
Acknowledgements
We would like to thank N. Bowen for helpful discussions and for providing many of the different protein preparations used in the in vitro MMR assays. This work was supported by NIH grants K99 ES026653 (E.M.G.), F32 CA210407 (W.J.G.) and R01 GM50006 (R.D.K.) and by the Ludwig Institute for Cancer Research (R.D.K. and C.D.P.).
Author information
Authors and Affiliations
Contributions
E.M.G., C.D.P. and R.D.K. conceived the overall experimental design; E.M.G. performed strain and plasmid construction, quantitative rate measurements and two-hybrid interaction analysis; C.M.R. aided in plasmid construction and performed Rad27 and Exo1 synthetic lethality experiments; W.J.G. performed MMR assays; B.-Z.L. performed Mlh1–Pms1 focus assays; C.D.P. analyzed SHIP box sequences and evolutionary relationships; E.M.G., C.D.P. and R.D.K. wrote the paper; and all of the authors revised and modified the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Yeast two-hybrid analysis of Exo1 C-terminal deletion mutants reveals two redundant sites are involved in the Msh2-Exo1 interaction.
a, Prey vectors bearing alleles of EXO1 were co-transformed with bait vectors bearing wild-type MSH2 to test for their ability to support the Msh2-Exo1 interaction. Vectors with full-length EXO1, the exo1F447A,F448A MIP-box-defective allele and the deletion constructs eliminating residues of Exo1 C terminal to amino acid 587 interacted with Msh2, as demonstrated by growth on CSM –Leu–Trp–His medium. Note that CSM –Leu–Trp medium is a growth control. In contrast, deletion mutants affecting amino acids N terminal to amino acid 587 disrupted Msh2 binding, as did the exo1Δ571–635,Δ671–702 allele. b, Prey vectors bearing EXO1 alleles were co-transformed with bait vectors bearing wild-type MLH1 to test for their ability to support the Mlh1-Exo1 interaction. All of the tested alleles of EXO1, except for the exo1F447A,F448A MIP-box-defective allele, encoded Exo1 variants that could bind to Mlh1, as demonstrated by growth on CSM –Leu–Trp–His medium.
Supplementary Figure 2 Patch test reveals that inactivation of both the MIP and SHIP box motifs in Exo1 cause a defect in Exo1-dependent MMR but not in the ability of Exo1 to suppress the synthetic lethality of exo1Δ and rad27Δ mutations.
a, Low-copy-number ARS-CEN plasmids without an insert or bearing different EXO1 constructs were tested for their ability to suppress the mutator phenotype of the exo1Δ pol30K217E double mutant (RDKY8077) or the exo1Δ pms1A99V double mutant (RDKY4192). Empty vector and vector bearing the nuclease-dead exo1D173A allele were unable to suppress the mutator phenotype, while the wild-type EXO1 substantially suppressed the increased mutation rate of the double mutant. An allele of EXO1 containing mutations that disrupted the MIP box (exo1F447A,F448A) and deleted the region including both SHIP boxes (exo1Δ571–702) was unable to suppress the mutator phenotype, equivalent to the empty vector or the nuclease-dead exo1D173A allele. All experiments were independently repeated a minimum of two times. b, An S. cerevisiae rad27Δ exo1Δ double-mutant strain containing a wild-type EXO1-bearing URA3 plasmid was transformed with TRP1 plasmids either without EXO1 or with various EXO1 mutations. The transformed strains were then plated either on YPD or CSM medium containing 5FOA to select for cells that lost the complementing EXO1-bearing URA3 plasmid. Neither the empty vector nor the nuclease-dead exo1D173A allele could suppress the synthetic lethality of the rad27Δ exo1Δ▯double mutations and grow on 5FOA-containing medium. In contrast, alleles of EXO1 containing mutations that disrupted the MIP box (exo1F447A,F448A) or deleted the region including both SHIP boxes (exo1Δ571–702) or lacked both functional MIP or SHIP boxes (exo1F447A,F448A,Δ571–702) would support growth on 5FOA-containing medium. Thus, defects in the MIP and SHIP boxes that cause MMR defects do not disrupt the functions of Exo1 required for survival of rad27Δ strains. All experiments were independently repeated a minimum of two times.
Supplementary Figure 3 Identification of putative SHIP box sequences in Exo1, Fun30, Dpb3, Bir1 and Utp18.
Left, 2D plot of each 7-mer peptide in the proteins plotted, with the SHIP peptide motif score generated with the PSSM along the x axis and the peptide disorder score generated using IUPRED (Bioinformatics 21, 3433–3434, 2005) along the y axis. Peptides with strong scores are labeled. Right, Diagram of the proteins with the position of the putative SHIP boxes displayed as black bars over a plot of the IUPRED long-range disorder score. Most putative SHIP boxes are present in extended unstructured regions at the N or C terminus of the proteins.
Supplementary Figure 4 Yeast two-hybrid analysis of the interaction between Msh2 and Utp18 and Bir1 and analysis of the levels of Pms1–4 × GFP foci in dpb3Δ fun30Δ single-mutant strains.
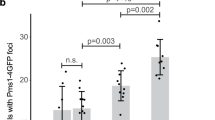
a, Prey vectors bearing either BIR1 or UTP18 were co-transformed with either empty bait vectors or bait vectors bearing wild-type MSH2, and interactions were evaluated as described in the legends to Fig. 1 and Supplementary Fig. 1. All experiments were independently repeated a minimum of four times. b, Pms1–4 × GFP foci were monitored in logarithmically growing asynchronous cultures by fluorescence microscopy, and the fraction of cells with one or more Pms1–4 × GFP foci was expressed as the fold change over wild type. The average value ( ± s.d.) from three independent experiments is presented.
Supplementary Figure 5 Conservation of SHIP and MIP box sequences in proteins from the Saccharomycetacae fungi.
The conservation of SHIP and MIP boxes present in S. cerevisiae was analyzed in key Saccharomycetacae fungi, including species that diverged before the whole-genome duplication that occurred during S. cerevisiae evolution (Yeast 24, 929–942, 2007). Presence of a MIP or SHIP box motif is indicated by “Y”; absence of the motif is indicated by “N”; and the absence of a homolog is indicated by “–”. The homologs of both EXO1 and DPB3 generated by the whole-genome duplication (ohnologs), DIN7 and DLS1, respectively, are also displayed when they exist. Phylogenic relationships were derived from a previous study (G3 6, 3927–3939, 2016).
Supplementary Figure 6 Phylogenetic distribution of the SHIP boxes in MCM9 and WDHD1/CTF4 in fungi, animals and closely related eukaryotes.
A “Y” in a black box indicates that the specific clade contains the SHIP box in the homologs. An “N” in a white box indicates that the specific clade contains the homologs but lacks an identifiable SHIP box. An asterisk next to the “Y” or “N” indicates that a very small number of species have a SHIP box status that differs from that of most of the species in the clade. A dash indicates that the clade lacks homologs of these genes, such as loss of MCM8 which is an MCM9 homolog, and MCM9 genes in the Dikarya fungi.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 and Supplementary Tables 1–4
Supplementary Dataset 1
Uncropped gel images
Supplementary Dataset 2
S. cerevisiae peptide PSSM/IUPRED scores
Supplementary Dataset 3
Human peptide PSSM/IUPRED scores
Supplementary Dataset 4
Assignment of the identities of fungal Exo1 homologs
Supplementary Dataset 5
Reannotation of the gene models for some eukaryotic Exo1 homologs
Rights and permissions
About this article
Cite this article
Goellner, E.M., Putnam, C.D., Graham, W.J. et al. Identification of Exo1-Msh2 interaction motifs in DNA mismatch repair and new Msh2-binding partners. Nat Struct Mol Biol 25, 650–659 (2018). https://doi.org/10.1038/s41594-018-0092-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-018-0092-y
This article is cited by
-
MicroRNA miR-21 Decreases Post-stroke Brain Damage in Rodents
Translational Stroke Research (2022)
-
Rad27 and Exo1 function in different excision pathways for mismatch repair in Saccharomyces cerevisiae
Nature Communications (2021)