Abstract
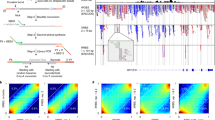
To identify oncogenes in leukemias, we performed large-scale resequencing of the leukemia genome using DNA sequence arrays that determine ∼9 Mbp of sequence corresponding to the exons or exon–intron boundaries of 5648 protein-coding genes. Hybridization of genomic DNA from CD34-positive blasts of acute myeloid leukemia (n=19) or myeloproliferative disorder (n=1) with the arrays identified 9148 nonsynonymous nucleotide changes. Subsequent analysis showed that most of these changes were also present in the genomic DNA of the paired controls, with 11 somatic changes identified only in the leukemic blasts. One of these latter changes results in a Met-to-Ile substitution at amino-acid position 511 of Janus kinase 3 (JAK3), and the JAK3(M511I) protein exhibited transforming potential both in vitro and in vivo. Further screening for JAK3 mutations showed novel and known transforming changes in a total of 9 out of 286 cases of leukemia. Our experiments also showed a somatic change responsible for an Arg-to-His substitution at amino-acid position 882 of DNA methyltransferase 3A, which resulted in a loss of DNA methylation activity of >50%. Our data have thus shown a unique profile of gene mutations in human leukemia.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Bentley DR, Balasubramanian S, Swerdlow HP, Smith GP, Milton J, Brown CG et al. (2008). Accurate whole human genome sequencing using reversible terminator chemistry. Nature 456: 53–59.
Byrd JC, Mrozek K, Dodge RK, Carroll AJ, Edwards CG, Arthur DC et al. (2002). Pretreatment cytogenetic abnormalities are predictive of induction success, cumulative incidence of relapse, and overall survival in adult patients with de novo acute myeloid leukemia: results from cancer and leukemia Group B (CALGB 8461). Blood 100: 4325–4336.
Choi YL, Kaneda R, Wada T, Fujiwara S, Soda M, Watanabe H et al. (2007). Identification of a constitutively active mutant of JAK3 by retroviral expression screening. Leuk Res 31: 203–209.
Ehrlich M . (2003). The ICF syndrome, a DNA methyltransferase 3B deficiency and immunodeficiency disease. Clin Immunol 109: 17–28.
El-Osta A . (2004). The rise and fall of genomic methylation in cancer. Leukemia 18: 233–237.
Greenberger JS, Sakakeeny MA, Humphries RK, Eaves CJ, Eckner RJ . (1983). Demonstration of permanent factor-dependent multipotential (erythroid/neutrophil/basophil) hematopoietic progenitor cell lines. Proc Natl Acad Sci USA 80: 2931–2935.
Greenman C, Stephens P, Smith R, Dalgliesh GL, Hunter C, Bignell G et al. (2007). Patterns of somatic mutation in human cancer genomes. Nature 446: 153–158.
Grimwade D, Walker H, Oliver F, Wheatley K, Harrison C, Harrison G et al. (1998). The importance of diagnostic cytogenetics on outcome in AML: analysis of 1612 patients entered into the MRC AML 10 trial. The Medical Research Council Adult and Children's Leukaemia Working Parties. Blood 92: 2322–2333.
Imanishi D, Miyazaki Y, Yamasaki R, Sawayama Y, Taguchi J, Tsushima H et al. (2007). Donor-derived DNA in fingernails among recipients of allogeneic hematopoietic stem-cell transplants. Blood 110: 2231–2234.
Iwama A, Oguro H, Negishi M, Kato Y, Morita Y, Tsukui H et al. (2004). Enhanced self-renewal of hematopoietic stem cells mediated by the polycomb gene product Bmi-1. Immunity 21: 843–851.
Jia D, Jurkowska RZ, Zhang X, Jeltsch A, Cheng X . (2007). Structure of Dnmt3a bound to Dnmt3L suggests a model for de novo DNA methylation. Nature 449: 248–251.
Kiyoi H, Yamaji S, Kojima S, Naoe T . (2007). JAK3 mutations occur in acute megakaryoblastic leukemia both in Down syndrome children and non-Down syndrome adults. Leukemia 21: 574–576.
Koinuma K, Kaneda R, Toyota M, Yamashita Y, Takada S, Choi YL et al. (2005). Screening for genomic fragments that are methylated specifically in colorectal carcinoma with a methylated MLH1 promoter. Carcinogenesis 26: 2078–2085.
Kralovics R, Passamonti F, Buser AS, Teo SS, Tiedt R, Passweg JR et al. (2005). A gain-of-function mutation of JAK2 in myeloproliferative disorders. N Engl J Med 352: 1779–1790.
Kubonishi I, Miyoshi I . (1983). Establishment of a Ph1 chromosome-positive cell line from chronic myelogenous leukemia in blast crisis. Int J Cell Cloning 1: 105–117.
Kwong YL, Wong KF, Chan V, Chan CH . (1996). Persistence of AML1 rearrangement in peripheral blood cells in t(8;21). Cancer Genet Cytogenet 88: 151–154.
Ley TJ, Mardis ER, Ding L, Fulton B, McLellan MD, Chen K et al. (2008). DNA sequencing of a cytogenetically normal acute myeloid leukaemia genome. Nature 456: 66–72.
Miyamoto T, Nagafuji K, Akashi K, Harada M, Kyo T, Akashi T et al. (1996). Persistence of multipotent progenitors expressing AML1/ETO transcripts in long-term remission patients with t(8;21) acute myelogenous leukemia. Blood 87: 4789–4796.
Miyamoto T, Weissman IL, Akashi K . (2000). AML1/ETO-expressing nonleukemic stem cells in acute myelogenous leukemia with 8;21 chromosomal translocation. Proc Natl Acad Sci USA 97: 7521–7526.
Nimer SD, Moore MA . (2004). Effects of the leukemia-associated AML1-ETO protein on hematopoietic stem and progenitor cells. Oncogene 23: 4249–4254.
Okano M, Bell DW, Haber DA, Li E . (1999). DNA methyltransferases Dnmt3a and Dnmt3b are essential for de novo methylation and mammalian development. Cell 99: 247–257.
Onishi M, Kinoshita S, Morikawa Y, Shibuya A, Phillips J, Lanier LL et al. (1996). Applications of retrovirus-mediated expression cloning. Exp Hematol 24: 324–329.
Patil N, Berno AJ, Hinds DA, Barrett WA, Doshi JM, Hacker CR et al. (2001). Blocks of limited haplotype diversity revealed by high-resolution scanning of human chromosome 21. Science 294: 1719–1723.
Pikman Y, Lee BH, Mercher T, McDowell E, Ebert BL, Gozo M et al. (2006). MPLW515L is a novel somatic activating mutation in myelofibrosis with myeloid metaplasia. PLoS Med 3: e270.
Russell SM, Tayebi N, Nakajima H, Riedy MC, Roberts JL, Aman MJ et al. (1995). Mutation of Jak3 in a patient with SCID: essential role of Jak3 in lymphoid development. Science 270: 797–800.
Sato T, Toki T, Kanezaki R, Xu G, Terui K, Kanegane H et al. (2008). Functional analysis of JAK3 mutations in transient myeloproliferative disorder and acute megakaryoblastic leukaemia accompanying Down syndrome. Br J Haematol 141: 681–688.
Schlenk RF, Dohner K, Krauter J, Frohling S, Corbacioglu A, Bullinger L et al. (2008). Mutations and treatment outcome in cytogenetically normal acute myeloid leukemia. N Engl J Med 358: 1909–1918.
Schwonzen M, Diehl V, Dellanna M, Staib P . (2007). Immunophenotyping of surface antigens in acute myeloid leukemia by flow cytometry after red blood cell lysis. Leuk Res 31: 113–116.
Shimada A, Taki T, Tabuchi K, Tawa A, Horibe K, Tsuchida M et al. (2006). KIT mutations, and not FLT3 internal tandem duplication, are strongly associated with a poor prognosis in pediatric acute myeloid leukemia with t(8;21): a study of the Japanese Childhood AML Cooperative Study Group. Blood 107: 1806–1809.
Sjoblom T, Jones S, Wood LD, Parsons DW, Lin J, Barber TD et al. (2006). The consensus coding sequences of human breast and colorectal cancers. Science 314: 268–274.
Suetake I, Miyazaki J, Murakami C, Takeshima H, Tajima S . (2003). Distinct enzymatic properties of recombinant mouse DNA methyltransferases Dnmt3a and Dnmt3b. J Biochem 133: 737–744.
Tallman MS, Altman JK . (2008). Curative strategies in acute promyelocytic leukemia. Hematol Am Soc Hematol Educ Program 2008: 391–399.
Walters DK, Mercher T, Gu TL, O'Hare T, Tyner JW, Loriaux M et al. (2006). Activating alleles of JAK3 in acute megakaryoblastic leukemia. Cancer Cell 10: 65–75.
Wheeler DA, Srinivasan M, Egholm M, Shen Y, Chen L, McGuire A et al. (2008). The complete genome of an individual by massively parallel DNA sequencing. Nature 452: 872–876.
Wong S, Witte ON . (2001). Modeling Philadelphia chromosome positive leukemias. Oncogene 20: 5644–5659.
Yamamoto Y, Kiyoi H, Nakano Y, Suzuki R, Kodera Y, Miyawaki S et al. (2001). Activating mutation of D835 within the activation loop of FLT3 in human hematologic malignancies. Blood 97: 2434–2439.
Acknowledgements
We thank D Cox, KA Frazer, DG Ballinger, J Montgomery, H Tao, C Chen, L Stuve, J Kwon, J Sheehan and Y Zhan for discussion on the wafer experiments, as well as JN Ihle, T Kitamura and SB Baylin for human JAK3 cDNA, the pMX plasmid and human DNMT3A cDNA, respectively. This study was supported in part by a grant for Third-Term Comprehensive Control Research for Cancer from the Ministry of Health, Labor, and Welfare of Japan, and by a grant for Scientific Research on Priority Areas ‘Applied Genomics’ from the Ministry of Education, Culture, Sports, Science, and Technology of Japan.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Oncogene website
Supplementary information
Rights and permissions
About this article
Cite this article
Yamashita, Y., Yuan, J., Suetake, I. et al. Array-based genomic resequencing of human leukemia. Oncogene 29, 3723–3731 (2010). https://doi.org/10.1038/onc.2010.117
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2010.117
Keywords
This article is cited by
-
Cancer Stem Cells are Actually Stem Cells with Disordered Differentiation: the Monophyletic Origin of Cancer
Stem Cell Reviews and Reports (2023)
-
Prevalence and dynamics of clonal hematopoiesis caused by leukemia-associated mutations in elderly individuals without hematologic disorders
Leukemia (2020)
-
Structural basis for impairment of DNA methylation by the DNMT3A R882H mutation
Nature Communications (2020)
-
Are peptides a solution for the treatment of hyperactivated JAK3 pathways?
Inflammopharmacology (2019)
-
Dynamics of DNMT3A mutation and prognostic relevance in patients with primary myelodysplastic syndrome
Clinical Epigenetics (2018)