Abstract
Neuronal inclusions of poly(GA), a protein unconventionally translated from G4C2 repeat expansions in C9ORF72, are abundant in patients with frontotemporal dementia (FTD) and amyotrophic lateral sclerosis (ALS) caused by this mutation. To investigate poly(GA) toxicity, we generated mice that exhibit poly(GA) pathology, neurodegeneration and behavioral abnormalities reminiscent of FTD and ALS. These phenotypes occurred in the absence of TDP-43 pathology and required poly(GA) aggregation. HR23 proteins involved in proteasomal degradation and proteins involved in nucleocytoplasmic transport were sequestered by poly(GA) in these mice. HR23A and HR23B similarly colocalized to poly(GA) inclusions in C9ORF72 expansion carriers. Sequestration was accompanied by an accumulation of ubiquitinated proteins and decreased xeroderma pigmentosum C (XPC) levels in mice, indicative of HR23A and HR23B dysfunction. Restoring HR23B levels attenuated poly(GA) aggregation and rescued poly(GA)-induced toxicity in neuronal cultures. These data demonstrate that sequestration and impairment of nuclear HR23 and nucleocytoplasmic transport proteins is an outcome of, and a contributor to, poly(GA) pathology.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
DeJesus-Hernandez, M. et al. Expanded GGGGCC hexanucleotide repeat in noncoding region of C9ORF72 causes chromosome 9p-linked FTD and ALS. Neuron 72, 245–256 (2011).
Renton, A.E. et al. A hexanucleotide repeat expansion in C9ORF72 is the cause of chromosome 9p21-linked ALS-FTD. Neuron 72, 257–268 (2011).
Belzil, V.V. et al. Reduced C9orf72 gene expression in c9FTD/ALS is caused by histone trimethylation, an epigenetic event detectable in blood. Acta Neuropathol. 126, 895–905 (2013).
Liu, E.Y. et al. C9orf72 hypermethylation protects against repeat expansion-associated pathology in ALS/FTD. Acta Neuropathol. 128, 525–541 (2014).
Xi, Z. et al. Hypermethylation of the CpG island near the G4C2 repeat in ALS with a C9orf72 expansion. Am. J. Hum. Genet. 92, 981–989 (2013).
van Blitterswijk, M. et al. Novel clinical associations with specific C9ORF72 transcripts in patients with repeat expansions in C9ORF72. Acta Neuropathol. 130, 863–876 (2015).
Gendron, T.F., Belzil, V.V., Zhang, Y.J. & Petrucelli, L. Mechanisms of toxicity in C9FTLD/ALS. Acta Neuropathol. 127, 359–376 (2014).
Sareen, D. et al. Targeting RNA foci in iPSC-derived motor neurons from ALS patients with a C9ORF72 repeat expansion. Sci. Transl. Med. 5, 208ra149 (2013).
Donnelly, C.J. et al. RNA toxicity from the ALS/FTD C9ORF72 expansion is mitigated by antisense intervention. Neuron 80, 415–428 (2013).
Cooper-Knock, J. et al. Sequestration of multiple RNA recognition motif-containing proteins by C9orf72 repeat expansions. Brain 137, 2040–2051 (2014).
Lee, Y.B. et al. Hexanucleotide repeats in ALS/FTD form length-dependent RNA foci, sequester RNA binding proteins, and are neurotoxic. Cell Rep. 5, 1178–1186 (2013).
Freibaum, B.D. et al. GGGGCC repeat expansion in C9orf72 compromises nucleocytoplasmic transport. Nature 525, 129–133 (2015).
Zhang, K. et al. The C9orf72 repeat expansion disrupts nucleocytoplasmic transport. Nature 525, 56–61 (2015).
Gendron, T.F. et al. Antisense transcripts of the expanded C9ORF72 hexanucleotide repeat form nuclear RNA foci and undergo repeat-associated non-ATG translation in c9FTD/ALS. Acta Neuropathol. 126, 829–844 (2013).
Ash, P.E. et al. Unconventional translation of C9ORF72 GGGGCC expansion generates insoluble polypeptides specific to c9FTD/ALS. Neuron 77, 639–646 (2013).
Mori, K. et al. Bidirectional transcripts of the expanded C9orf72 hexanucleotide repeat are translated into aggregating dipeptide repeat proteins. Acta Neuropathol. 126, 881–893 (2013).
Mori, K. et al. The C9orf72 GGGGCC repeat is translated into aggregating dipeptide-repeat proteins in FTLD/ALS. Science 339, 1335–1338 (2013).
Zu, T. et al. RAN proteins and RNA foci from antisense transcripts in C9ORF72 ALS and frontotemporal dementia. Proc. Natl. Acad. Sci. USA 110, E4968–E4977 (2013).
Jovičić, A. et al. Modifiers of C9orf72 dipeptide repeat toxicity connect nucleocytoplasmic transport defects to FTD/ALS. Nat. Neurosci. 18, 1226–1229 (2015).
Yamakawa, M. et al. Characterization of the dipeptide repeat protein in the molecular pathogenesis of c9FTD/ALS. Hum. Mol. Genet. 24, 1630–1645 (2014).
Mizielinska, S. et al. C9orf72 repeat expansions cause neurodegeneration in Drosophila through arginine-rich proteins. Science 345, 1192–1194 (2014).
May, S. et al. C9orf72 FTLD/ALS-associated Gly-Ala dipeptide repeat proteins cause neuronal toxicity and Unc119 sequestration. Acta Neuropathol. 128, 485–503 (2014).
Wen, X. et al. Antisense proline-arginine RAN dipeptides linked to C9ORF72-ALS/FTD form toxic nuclear aggregates that initiate in vitro and in vivo neuronal death. Neuron 84, 1213–1225 (2014).
Kwon, I. et al. Poly-dipeptides encoded by the C9orf72 repeats bind nucleoli, impede RNA biogenesis, and kill cells. Science 345, 1139–1145 (2014).
Zhang, Y.J. et al. Aggregation-prone c9FTD/ALS poly(GA) RAN-translated proteins cause neurotoxicity by inducing ER stress. Acta Neuropathol. 128, 505–524 (2014).
Tao, Z. et al. Nucleolar stress and impaired stress granule formation contribute to C9orf72 RAN translation-induced cytotoxicity. Hum. Mol. Genet. 24, 2426–2441 (2015).
Mackenzie, I.R. et al. Quantitative analysis and clinico-pathological correlations of different dipeptide repeat protein pathologies in C9ORF72 mutation carriers. Acta Neuropathol. 130, 845–861 (2015).
Schludi, M.H. et al. Distribution of dipeptide repeat proteins in cellular models and C9orf72 mutation cases suggests link to transcriptional silencing. Acta Neuropathol. 130, 537–555 (2015).
Woerner, A.C. et al. Cytoplasmic protein aggregates interfere with nucleo-cytoplasmic transport of protein and RNA. Science 351, 173–176 (2015).
Fecto, F. et al. SQSTM1 mutations in familial and sporadic amyotrophic lateral sclerosis. Arch. Neurol. 68, 1440–1446 (2011).
Le Ber, I. et al. French Clinical and Genetic Research Network on FTD/FTD-ALS. SQSTM1 mutations in French patients with frontotemporal dementia or frontotemporal dementia with amyotrophic lateral sclerosis. JAMA Neurol. 70, 1403–1410 (2013).
Deng, H.X. et al. Mutations in UBQLN2 cause dominant X-linked juvenile and adult-onset ALS and ALS/dementia. Nature 477, 211–215 (2011).
Dougan, L., Li, J., Badilla, C.L., Berne, B.J. & Fernandez, J.M. Single homopolypeptide chains collapse into mechanically rigid conformations. Proc. Natl. Acad. Sci. USA 106, 12605–12610 (2009).
Popiel, H.A. et al. Disruption of the toxic conformation of the expanded polyglutamine stretch leads to suppression of aggregate formation and cytotoxicity. Biochem. Biophys. Res. Commun. 317, 1200–1206 (2004).
Pelassa, I. et al. Association of polyalanine and polyglutamine coiled coils mediates expansion disease-related protein aggregation and dysfunction. Hum. Mol. Genet. 23, 3402–3420 (2014).
Chiba, T. et al. Amyloid fibril formation in the context of full-length protein: effects of proline mutations on the amyloid fibril formation of beta2-microglobulin. J. Biol. Chem. 278, 47016–47024 (2003).
Mocanu, M.M. et al. The potential for beta-structure in the repeat domain of tau protein determines aggregation, synaptic decay, neuronal loss, and coassembly with endogenous Tau in inducible mouse models of tauopathy. J. Neurosci. 28, 737–748 (2008).
Dantuma, N.P., Heinen, C. & Hoogstraten, D. The ubiquitin receptor Rad23: at the crossroads of nucleotide excision repair and proteasomal degradation. DNA Repair (Amst.) 8, 449–460 (2009).
Chew, J. et al. Neurodegeneration. C9ORF72 repeat expansions in mice cause TDP-43 pathology, neuronal loss, and behavioral deficits. Science 348, 1151–1154 (2015).
Mitchell, J.M., Mansfeld, J., Capitanio, J., Kutay, U. & Wozniak, R.W. Pom121 links two essential subcomplexes of the nuclear pore complex core to the membrane. J. Cell Biol. 191, 505–521 (2010).
Antonin, W., Franz, C., Haselmann, U., Antony, C. & Mattaj, I.W. The integral membrane nucleoporin pom121 functionally links nuclear pore complex assembly and nuclear envelope formation. Mol. Cell 17, 83–92 (2005).
Ng, J.M. et al. A novel regulation mechanism of DNA repair by damage-induced and RAD23-dependent stabilization of xeroderma pigmentosum group C protein. Genes Dev. 17, 1630–1645 (2003).
Melis, J.P.M., Luijten, M., Mullenders, L.H.F. & van Steeg, H. The role of XPC: implications in cancer and oxidative DNA damage. Mutat. Res. 728, 107–117 (2011).
Zhang, Y., Rohde, L.H. & Wu, H. Involvement of nucleotide excision and mismatch repair mechanisms in double strand break repair. Curr. Genomics 10, 250–258 (2009).
Ng, J.M. et al. Developmental defects and male sterility in mice lacking the ubiquitin-like DNA repair gene mHR23B. Mol. Cell. Biol. 22, 1233–1245 (2002).
Davidson, Y. et al. Neurodegeneration in frontotemporal lobar degeneration and motor neurone disease associated with expansions in C9orf72 is linked to TDP-43 pathology and not associated with aggregated forms of dipeptide repeat proteins. Neuropathol. Appl. Neurobiol. Published online 5 November 2015 (doi:10.1111/nan.12292).
Todd, T.W. & Lim, J. Aggregation formation in the polyglutamine diseases: protection at a cost? Mol. Cells 36, 185–194 (2013).
Bergink, S. et al. The DNA repair-ubiquitin-associated HR23 proteins are constituents of neuronal inclusions in specific neurodegenerative disorders without hampering DNA repair. Neurobiol. Dis. 23, 708–716 (2006).
Chakrabarty, P. et al. Capsid serotype and timing of injection determines AAV transduction in the neonatal mice brain. PLoS One 8, e67680 (2013).
Kim, J.Y. et al. Viral transduction of the neonatal brain delivers controllable genetic mosaicism for visualising and manipulating neuronal circuits in vivo. Eur. J. Neurosci. 37, 1203–1220 (2013).
Xu, Y.F. et al. Wild-type human TDP-43 expression causes TDP-43 phosphorylation, mitochondrial aggregation, motor deficits, and early mortality in transgenic mice. J. Neurosci. 30, 10851–10859 (2010).
Lin, W.L., Dickson, D.W. & Sahara, N. Immunoelectron microscopic and biochemical studies of caspase-cleaved tau in a mouse model of tauopathy. J. Neuropathol. Exp. Neurol. 70, 779–787 (2011).
Murray, M.E. et al. A quantitative postmortem MRI design sensitive to white matter hyperintensity differences and their relationship with underlying pathology. J. Neuropathol. Exp. Neurol. 71, 1113–1122 (2012).
Shinohara, M., Petersen, R.C., Dickson, D.W. & Bu, G. Brain regional correlation of amyloid-β with synapses and apolipoprotein E in non-demented individuals: potential mechanisms underlying regional vulnerability to amyloid-β accumulation. Acta Neuropathol. 125, 535–547 (2013).
Zhang, Y.J. et al. Aberrant cleavage of TDP-43 enhances aggregation and cellular toxicity. Proc. Natl. Acad. Sci. USA 106, 7607–7612 (2009).
Gendron, T.F. et al. Cerebellar c9RAN proteins associate with clinical and neuropathological characteristics of C9ORF72 repeat expansion carriers. Acta Neuropathol. 130, 559–573 (2015).
Cook, C. et al. Severe amygdala dysfunction in a MAPT transgenic mouse model of frontotemporal dementia. Neurobiol. Aging 35, 1769–1777 (2014).
Acknowledgements
We are grateful to all patients who agreed to donate post-mortem tissue. This work was supported by the US National Institutes of Health (NIH) National Institute on Aging (R01AG026251 (L.P.)); NIH National Institute of Neurological Disorders and Stroke (R21NS079807 (Y.-J.Z. and J.D.F.); R21NS089979 (T.F.G. and K.B.B.); F32NS087842 (J.J.); R01NS080882 (R.R.); R01NS085207 (J.D.R.); U54NS091046 (J.D.R.); R01NS063964 (L.P.); R01NS077402 (L.P.), R21NS084528 (L.P.); P01NS084974 (L.P., D.D., K.B.B. and R.R.); R01NS088689 (L.P.)); National Institute of Environmental Health Services (R01ES20395 (L.P.); Department of Defense (ALSRP AL130125 (L.P.)); Mayo Clinic Foundation (L.P.); Mayo Clinic Center for Individualized Medicine (L.P. and K.B.B.); Alzheimer's Association (NIRP-14-304425 (Y.-J.Z.); NIRP-12-259289 (J.D.F.)); Amyotrophic Lateral Sclerosis Association (Y.-J.Z., T.F.G., K.B.B., D.W.C. and L.P.); Robert Packard Center for ALS Research at Johns Hopkins (J.D.R. and L.P.), Target ALS (C.L.-T., J.D.R. and L.P.); Brain Science Institute (J.D.R.); the Ludwig Institute for Cancer Research (D.W.C. and C.L.T.), and the European Union's Seventh Framework Programme (FP7/2014-2019 grant 617198 (D.E.)). J.C.G. is the recipient of a National Science Foundation Graduate Research Fellowship Award, a Thomas Shortman Training Fund Graduate Scholarship and an Axol Science Scholarship.
Author information
Authors and Affiliations
Contributions
L.P. and Y.-J.Z. contributed to the conception and design. Y.-J.Z. performed immunoblots, quantitative reverse-transcription PCR (qRT-PCR), co-immunoprecipitation and behavioral tests; T.F.G. completed anti-GA antibody generation and characterization, and performed poly(GA) assays with L.D.; J.C.G. and J.D.R. performed immunofluorescence staining for RanGAP1 and Pom121. H.S. performed ICV injection and behavioral tests; Y.-F.X. performed silver staining, immunofluorescence staining and primary neuronal cultures; Y.-F.X. and Z.S.W. quantified the Purkinje cells in cerebellum. M.E.M. and A.M.L. quantified neuronal loss and gliosis burden; M.S. and G.B. contributed to ELISA; W.-L.L. performed immunoEM; J.G. and A.G. performed immunofluorescence staining and immunoblotting; J.N.S. prepared primary neurons; K.J.-W. made plasmids; J.T. and M.Y. harvested mice and prepared brain lysates; E.A.P. produced AAV1; J.C. aided with ICV injections; M.C.-C. performed immunohistochemistry staining; A.K. and J.D.F. contributed to behavioral tests; J.D.B. and C.A.D. contributed to the purification of recombinant protein and the transmission electron microscopy study; J.J., C.L.-T., D.E. and D.W.C. characterized and provided anti-poly(GA) antibodies. R.R., K.B.B. and D.W.D. contributed to the tissue collection. C.D.L. analyzed data. L.P., Y.-J.Z., T.F.G. and R.B.K. analyzed data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Neuropathology of poly(GA) inclusions in the brains of GFP-(GA)50 mice.
Immunohistochemical analysis in the indicated brain regions of GFP-(GA)50 mice with (a) anti-GFP antibody or (b) anti-GA antibody. Scale bars, 20 μm. (c) Double immunofluorescence staining in the cortex of GFP-(GA)50 mice for the indicated proteins. The arrows point to MAP2-positive neurons and the arrowhead to an astrocyte. Scale bars, 10 μm. (d) Immunohistochemical analysis in the cortex and hippocampus of GFP, GFP-(GA)50 and GFP-(GA)50-mut mice with an anti-ubiquitin antibody. Scale bars, 20 μm.
Supplementary Figure 2 HR23, RanGAP1 and Pom121 form inclusions in the brains of GFP-(GA)50 and (G4C2)66 mice.
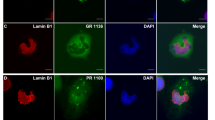
(a) Immunohistochemical analysis of HR23A and HR23B proteins in the hippocampus of 6 month-old GFP, GFP-(GA)50 and GFP-(GA)50-mut mice. Scale bars, 20 μm. (b) Double immunofluorescence staining for GFP-(GA)50 and HR23A or HR23B in the hippocampus of 6 month-old GFP-(GA)50 mice. Scale bars, 5 μm. (c) Immunofluorescence staining for HR23B in the cortex of 6 month-old (G4C2)66 mice. Scale bar, 10 μm. (d) Representative images of immunohistochemical analysis of HR23A and HR23B in the hippocampus of c9FTD/ALS subjects (n = 7) or healthy controls (n = 4). Scale bars, 20 μm. (e) Double immunofluorescence staining for RanGAP1 and either HR23A or HR23B in the cortex of 6 month-old GFP-(GA)50 mice. Scale bars, 10 μm. (f) Immunofluorescence staining for RanGAP1 or Pom121 in the cortex of 6 month-old (G4C2)66 mice. Scale bars, 10 μm.
Supplementary Figure 3 HR23 proteins interact with, and are sequestered by, poly(GA) proteins.
(a) Cytoplasmic and nuclear fractions were prepared from HEK293T cells exogenously expressing GFP, GFP-(GA)50 or GFP-(GA)50-mut, followed by immunoblots analysis using the indicated antibodies. Tubulin and Lamin A/C were used as cytoplasmic and nuclear markers, respectively. (b) Protein complexes were immunoprecipitated from the indicated input lysates (top left) from HEK293T cells exogenously expressing GFP, GFP-(GA)50 or GFP-(GA)50-mut with antibodies to GFP, HR23A or HR23B, followed by immunoblot analysis using the indicated antibodies. (c) Protein complexes were immunoprecipitated from the indicated input lysates (left, top and bottom) from HEK293T cells exogenously expressing GFP, GFP-(GA)50, GFP-(GR)50, or GFP-(GP)47 with an antibody to HR23B, followed by immunoblot analysis using antibodies to GFP and poly(GR). (d) Double immunofluorescence staining for HR23B and poly(GA), poly(GP) or poly(GR) in brains of 6 month-old (G4C2)66 mice. Scale bar, 5 μm. (e) Double immunofluorescence staining for HR23B and poly(GA), poly(GP) or poly(GR) in the hippocampus of c9FTD/ALS patients. Scale bar, 5 μm. For a-c, full-length immunoblots are presented in Supplementary Figure 10.
Supplementary Figure 4 XPC is sequestered into poly(GA) inclusions in the hippocampus of GFP-(GA)50 mice.
(a) Immunohistochemical analysis of XPC in the hippocampus of GFP, GFP-(GA)50 and GFP-(GA)50-mut mice. Scale bar, 20 μm. (b) Double immunofluorescence staining of XPC and poly(GA) proteins in the hippocampus of GFP-(GA)50 mice. Scale bar, 5 μm.
Supplementary Figure 5 Analysis of brain morphology, body weight and motor cortex neurons in GFP-(GA)50 mice.
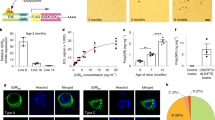
(a) Gross morphological analysis with hematoxylin and eosin staining of brains from 6 month-old GFP, GFP-(GA)50 and GFP-(GA)50-mut mice. Scale bar, 5 mm. (b) The mean body weight of 6 month-old male and female GFP, GFP-(GA)50 and GFP-(GA)50-mut mice, using 6–8 male mice or 4 female mice per group. Data are presented as mean ± s.e.m. Male mice: P < 0.0001, as analyzed by one-way ANOVA; ****P < 0.0001 and ***P = 0.0002, Tukey’s post-hoc analysis. Female mice: P = 0.0724, one-way ANOVA. n.s., not significant. (c) Immunohistochemical analysis of NeuN in layer V of the motor cortex of GFP, GFP-(GA)50 and GFP-(GA)50-mut mice. Scale bar, 30 μm.
Supplementary Figure 6 Poly(GA) inclusions in 4- to 6-week-old GFP-(GA)50 mice.
Immunohistochemical analysis of cortex and hippocampus of GFP, GFP-(GA)50 and GFP-(GA)50-mut mice with (a) anti-GFP antibody or (b) anti-ubiquitin antibody. Scale bars, 20 μm.
Supplementary Figure 7 No signs of neurodegeneration were observed in 4- to 6-week-old GFP-(GA)50 mice.
(a) Graph showing the mean brain weight of mice expressing GFP, GFP-(GA)50 or GFP-(GA)50-mut (n = 5–7 per group). (b) The mean body weight of male and female GFP, GFP-(GA)50 and GFP-(GA)50-mut mice using 2–5 male mice or 1–3 female mice. (c) Representative images of NeuN-labeled cells in the motor cortex and hippocampus of GFP, GFP-(GA)50 or GFP-(GA)50-mut mice. Scale bars, 200 μm. (d) Quantification of the number of NeuN-positive cells in the cortex (left) or motor cortex (right) of GFP, GFP-(GA)50 or GFP-(GA)50-mut mice (n = 5–7 per group). (e) Quantification of the number of Purkinje cells in the cerebellum of GFP, GFP-(GA)50 or GFP-(GA)50-mut mice (n = 5–7 per group). (f) Representative images of GFAP staining to identify reactive astrocytes in the motor cortex and hippocampus of GFP, GFP-(GA)50 or GFP-(GA)50-mut mice. Scale bars, 100 μm. Data are presented as mean ± s.e.m., and analyzed by one-way ANOVA; P = 0.0988 (a), P = 0.7026 (b), P = 0.0563 (d, Cortex), P = 0.1609 (d, Motor cortex) and P = 0.9042 (e). n.s., not significant.
Supplementary Figure 8 Exogenous HR23B does not decrease poly(GR) levels nor attenuate poly(GR)-induced neurotoxicity.
(a) Immunoblot and (b) densitometric analysis of immunoblots for the indicated proteins to determine their levels of expression in primary neurons transduced to express GFP-(GR)50 or GFP in the presence or absence of exogenous Myc-tagged HR23B. Data are presented as mean ± s.e.m. from 3 separate experiments. In b, left: P = 0.0005, one-way ANOVA; P = 0.6252 (GFP-(GR)50+Vector vs. GFP-(GR)50+HR23B-Myc), Tukey’s post-hoc analysis. Right: P < 0.0001, one-way ANOVA; ****P < 0.0001 and P = 0.7798 (GFP-(GR)50+Vector vs. GFP-(GR)50+HR23B-Myc), Tukey’s post-hoc analysis. n.s., not significant. For a, full-length immunoblots are presented in Supplementary Figure 10.
Supplementary Figure 9 Full-length immunoblots for main figures.
The region delineated by the box on each blot is the image shown in the corresponding figure.
Supplementary Figure 10 Full-length immunoblots for supplementary figures.
The region delineated by the box on each blot is the image shown in the corresponding figure.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10 and Supplementary Tables 1–3 (PDF 1548 kb)
Rights and permissions
About this article
Cite this article
Zhang, YJ., Gendron, T., Grima, J. et al. C9ORF72 poly(GA) aggregates sequester and impair HR23 and nucleocytoplasmic transport proteins. Nat Neurosci 19, 668–677 (2016). https://doi.org/10.1038/nn.4272
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn.4272
This article is cited by
-
Nuclear-import receptors as gatekeepers of pathological phase transitions in ALS/FTD
Molecular Neurodegeneration (2024)
-
Innate immune sensing of lysosomal dysfunction drives multiple lysosomal storage disorders
Nature Cell Biology (2024)
-
Repeat length of C9orf72-associated glycine–alanine polypeptides affects their toxicity
Acta Neuropathologica Communications (2023)
-
Reconstitution of C9orf72 GGGGCC repeat-associated non-AUG translation with purified human translation factors
Scientific Reports (2023)
-
Proximity proteomics of C9orf72 dipeptide repeat proteins identifies molecular chaperones as modifiers of poly-GA aggregation
Acta Neuropathologica Communications (2022)