Abstract
The transcription regulator YAP controls organ size by regulating cell growth, proliferation and apoptosis. However, whether YAP has a role in innate antiviral immunity is largely unknown. Here we found that YAP negatively regulated an antiviral immune response. YAP deficiency resulted in enhanced innate immunity, a diminished viral load, and morbidity in vivo. YAP blocked dimerization of the transcription factor IRF3 and impeded translocation of IRF3 to the nucleus after viral infection. Notably, virus-activated kinase IKKɛ phosphorylated YAP at Ser403 and thereby triggered degradation of YAP in lysosomes and, consequently, relief of YAP-mediated inhibition of the cellular antiviral response. These findings not only establish YAP as a modulator of the activation of IRF3 but also identify a previously unknown regulatory mechanism independent of the kinases Hippo and LATS via which YAP is controlled by the innate immune pathway.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Change history
20 July 2017
In the version of this article initially published, the source of the HSV-1 virus stock used in the study was identified incorrectly as a purchase from the Wuhan Institute of Virology, Chinese Academy of Sciences. The correct Acknowledgements section should begin "We thank F. Zhao (Wuhan Institute of Virology, Chinese Academy of Sciences) for HSV-1" and the second sentence of the third subsection of Online Methods ('Cells and reagents') should read "Herpes simplex virus type 1 (HSV-1) was obtained from F. Zhao (Wuhan Institute of Virology, Chinese Academy of Sciences)." The error has been corrected in the HTML and PDF versions of the article.
18 October 2017
Nat. Immunol. 18, 733–743 (2017); published online 8 May 2017; corrected after print 20 July 2017 In the version of this article initially published, the source of the HSV-1 virus stock used in the study was identified incorrectly as a purchase from the Wuhan Institute of Virology, Chinese Academy of Sciences.
References
Alexopoulou, L., Holt, A.C., Medzhitov, R. & Flavell, R.A. Recognition of double-stranded RNA and activation of NF-κB by Toll-like receptor 3. Nature 413, 732–738 (2001).
Wu, J. & Chen, Z.J. Innate immune sensing and signaling of cytosolic nucleic acids. Annu. Rev. Immunol. 32, 461–488 (2014).
Kato, H., Takahasi, K. & Fujita, T. RIG-I-like receptors: cytoplasmic sensors for non-self RNA. Immunol. Rev. 243, 91–98 (2011).
Sun, L., Wu, J., Du, F., Chen, X. & Chen, Z.J. Cyclic GMP-AMP synthase is a cytosolic DNA sensor that activates the type I interferon pathway. Science 339, 786–791 (2013).
Sato, S. et al. Toll/IL-1 receptor domain-containing adaptor inducing IFN-β (TRIF) associates with TNF receptor-associated factor 6 and TANK-binding kinase 1, and activates two distinct transcription factors, NF-kappa B and IFN-regulatory factor-3, in the Toll-like receptor signaling. J. Immunol. 171, 4304–4310 (2003).
Seth, R.B., Sun, L., Ea, C.K. & Chen, Z.J. Identification and characterization of MAVS, a mitochondrial antiviral signaling protein that activates NF-κB and IRF 3. Cell 122, 669–682 (2005).
Ishikawa, H. & Barber, G.N. STING is an endoplasmic reticulum adaptor that facilitates innate immune signalling. Nature 455, 674–678 (2008).
Takeuchi, O. & Akira, S. Pattern recognition receptors and inflammation. Cell 140, 805–820 (2010).
Kawai, T. & Akira, S. The role of pattern-recognition receptors in innate immunity: update on Toll-like receptors. Nat. Immunol. 11, 373–384 (2010).
Takeuchi, O. & Akira, S. Innate immunity to virus infection. Immunol. Rev. 227, 75–86 (2009).
Kawasaki, T., Kawai, T. & Akira, S. Recognition of nucleic acids by pattern-recognition receptors and its relevance in autoimmunity. Immunol. Rev. 243, 61–73 (2011).
Cao, X. Self-regulation and cross-regulation of pattern-recognition receptor signalling in health and disease. Nat. Rev. Immunol. 16, 35–50 (2016).
Moroishi, T., Hansen, C.G. & Guan, K.L. The emerging roles of YAP and TAZ in cancer. Nat. Rev. Cancer 15, 73–79 (2015).
Yu, F.X., Zhao, B. & Guan, K.L. Hippo pathway in organ size control, tissue homeostasis, and cancer. Cell 163, 811–828 (2015).
Meng, Z., Moroishi, T. & Guan, K.L. Mechanisms of Hippo pathway regulation. Genes Dev. 30, 1–17 (2016).
Rauskolb, C., Sun, S., Sun, G., Pan, Y. & Irvine, K.D. Cytoskeletal tension inhibits Hippo signaling through an Ajuba-Warts complex. Cell 158, 143–156 (2014).
Wang, L. et al. Integrin-YAP/TAZ-JNK cascade mediates atheroprotective effect of unidirectional shear flow. Nature 540, 579–582 (2016).
Yu, F.X. et al. Regulation of the Hippo-YAP pathway by G-protein-coupled receptor signaling. Cell 150, 780–791 (2012).
Mo, J.S. et al. Cellular energy stress induces AMPK-mediated regulation of YAP and the Hippo pathway. Nat. Cell Biol. 17, 500–510 (2015).
Santinon, G., Pocaterra, A. & Dupont, S. Control of YAP/TAZ activity by metabolic and nutrient-sensing pathways. Trends Cell Biol. 26, 289–299 (2016).
Morin-Kensicki, E.M. et al. Defects in yolk sac vasculogenesis, chorioallantoic fusion, and embryonic axis elongation in mice with targeted disruption of Yap65. Mol. Cell. Biol. 26, 77–87 (2006).
Dragan, A.I., Hargreaves, V.V., Makeyeva, E.N. & Privalov, P.L. Mechanisms of activation of interferon regulator factor 3: the role of C-terminal domain phosphorylation in IRF-3 dimerization and DNA binding. Nucleic Acids Res. 35, 3525–3534 (2007).
Panne, D., McWhirter, S.M., Maniatis, T. & Harrison, S.C. Interferon regulatory factor 3 is regulated by a dual phosphorylation-dependent switch. J. Biol. Chem. 282, 22816–22822 (2007).
Lin, R., Mamane, Y. & Hiscott, J. Structural and functional analysis of interferon regulatory factor 3: localization of the transactivation and autoinhibitory domains. Mol. Cell. Biol. 19, 2465–2474 (1999).
Zhu, M., Fang, T., Li, S., Meng, K. & Guo, D. Bipartite nuclear localization signal controls nuclear import and DNA-binding activity of IFN regulatory factor 3. J. Immunol. 195, 289–297 (2015).
Kutay, U., Izaurralde, E., Bischoff, F.R., Mattaj, I.W. & Görlich, D. Dominant-negative mutants of importin-beta block multiple pathways of import and export through the nuclear pore complex. EMBO J. 16, 1153–1163 (1997).
Marfori, M. et al. Molecular basis for specificity of nuclear import and prediction of nuclear localization. Biochim. Biophys. Acta 1813, 1562–1577 (2011).
Zhao, B. et al. Inactivation of YAP oncoprotein by the Hippo pathway is involved in cell contact inhibition and tissue growth control. Genes Dev. 21, 2747–2761 (2007).
Boro, M., Singh, V. & Balaji, K.N. Mycobacterium tuberculosis-triggered Hippo pathway orchestrates CXCL1/2 expression to modulate host immune responses. Sci. Rep. 6, 37695 (2016).
Meng, F. et al. Mst1 shuts off cytosolic antiviral defense through IRF3 phosphorylation. Genes Dev. 30, 1086–1100 (2016).
Liu, B. et al. Toll receptor-mediated hippo signaling controls innate immunity in Drosophila. Cell 164, 406–419 (2016).
Zhang, N. et al. The Merlin/NF2 tumor suppressor functions through the YAP oncoprotein to regulate tissue homeostasis in mammals. Dev. Cell 19, 27–38 (2010).
Ran, F.A. et al. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Acknowledgements
We thank F. Zhao (Wuhan Institute of Virology, Chinese Academy of Sciences) for HSV-1; D. Pan (Johns Hopkins University School of Medicine) for Yap1fl/fl mice; M. Rabelink (Leiden University Medical Center) for shRNA constructs; P. ten Dijke for help with writing; and J. Dai for discussion. Supported by a special program from Ministry of Science and Technology of China (2016YFA0502500 to L.Z.), the Chinese National Natural Science Funds (31571460 to F.Z., 31471315 and 31671457 to L.Z.), PCSIRT (IRT1075 to X.G.) and Jiangsu National Science Foundation (BK20150354 to F.Z.).
Author information
Authors and Affiliations
Contributions
S.W., L.Z. and F.Z. designed the experiments and analyzed the data; S.W., F.X., F.C., Z.Z., and T.D. performed the experiments; B.Y., L.G., L.W., L.L., J. Jia and J. Jin contributed to writing, discussions and agreement with the conclusions presented; and H.v.D. L.Z. and F.Z. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 YAP negatively regulates IFN-β signaling.
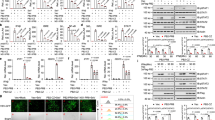
(a) Immunoblot analysis of Yap knockdown efficiency with sh-Yap (#1 to #4 independent constructs) in Raw264.7 cells. (b) IFN-β-Luc and PRDs I-III-Luc activity in Raw264.7 cells depleted for Yap and stimulated with SeV for 12 h. Results are shown as mean + sd of triplicates of at least two independent experiments. *P < 0.05 (two-tailed Student's t-test.) (c) qPCR analysis of sh-YAP #1 efficiency (left panel) and IFNB1 mRNA level (right panel) in control and YAP depleted THP1 cells followed by SeV infection at the indicated time points. (d) IFN-β-Luc and PRDs I-III-Luc activity in HEK293T cells transfected with respective reporters and YAP2 or YAP4 expression plasmids followed by stimulation with SeV for 12 h. (e) qPCR analysis of HEK293T cells transfected with empty vector (Co.vec) and YAP2 or YAP4 expression plasmids for 48 h followed by Sev infection (left) or Poly (I:C) transfection at the indicated time points. All qPCR results are shown as mean + sd of triplicates of at least two independent experiments. *P < 0.05 and **P < 0.01 (two-tailed Student's t-test.) (f) Fluorescence microscopy of VSV–GFP levels in HEK293T cells transfected with YAP2 or YAP4 expression plasmids followed by infection for 12 h with VSV–GFP (MOI 0.1) (bright-field, upper; fluorescence, bottom). Scale bars, 100 μm. Representative results are shown of least two independent experiments.
Supplementary Figure 2 YAP deficiency potentiates IFN-β signaling.
(a, b) qPCR analysis of Ifnb1, Ccl5 and Cxcl10 mRNA levels in wild type and Yap1+/- mouse bone marrow derived macrophages (BMDMs) infected with SeV (a) or stimulated with 5'-pppRNA for the indicated time points. (c, d) qPCR analysis of Ifnb1 levels in wild type and Yap1+/- BMDMs transfected with Poly (I:C) (c) or infected with HSV-1 (MOI, 10) (d) for the indicated time points.(e, f) qPCR analysis of Ifnb1, ccl5 and Cxcl10 mRNA levels in wild type and Yap1+/- MEFs infected with SeV (e) or stimulated with 5'-ppp RNA (f) for the indicated time points. (g, h) qPCR analysis of Ifnb1 levels in wild type and Yap1+/- MEFs transfected with Poly (I:C) (g) or infected with HSV-1 (MOI, 10) (h) for the indicated time points. All qPCR results are shown as mean + sd of triplicates of at least two independent experiments. *P < 0.05 and **P < 0.01 (two-tailed Student's t-test.) (i) Survival of Yap1+/+ and Yap1+/− mice (n=10 for each group) infected intraperitoneally with VSV (5×108 PFU per mouse). *P < 0.05 (two-way analysis of variance (ANOVA)). (j) Representative hematoxylin-and-eosin-stained images of lung sections from mice as in a. Scale bars, 100 μm. (k) HSV-1 viral titers in brain of Yap WT and heterogeneous mice (n = 6 mice per group) infected with HSV-1 for 72 h. (l) Survival of Yap1+/- and WT mice (n = 10 mice per group) after intravenous injection of HSV-1 (2×107 PFU per mouse). *P < 0.05, *** P < 0.01 (two-tailed Student's t-test in k or two-way analysis of variance (ANOVA) in l).
Supplementary Figure 3 IFN-β signaling is upregulated in YAP-deficient macrophages.
qPCR of Ifnb1, Cxcl10 and Ccl5 mRNA levels in Yap1fl/fl Lyz2-Cre+ and Yap1fl/fl Lyz2-Cre- peritoneal macrophages infected or stimulated with SeV, 5'-pppRNA, VSV(MOI=1) or HSV-1 (MOI=0.1). All qPCR results are shown as mean + sd of triplicates of at least two independent experiments. *P < 0.05 and **P < 0.01 (two-tailed Student's t-test.)
Supplementary Figure 4 YAP retains IRF3 in the cytoplasm.
(a) IFN-β-Luc activity (left) and IFNB1 mRNA in HEK293T cells depleted with YAP and transfected with cGAS+STING, RIG-I N-terminal 2CARD (RIG-IN), MAVS, TBK1, IKKɛ or IRF3-5D expression plasmids as indicated. Results are shown as mean + sd of triplicates of at least two independent experiments. *P < 0.05 (two-tailed Student's t-test.) (b) Immunoblot (IB) of total cell lysate (TCL) and immunoprecipitate derived from HEK293T cells transfected with Myc-IRF3 and Flag-YAP1 or 2 expression plasmids as indicated. (c) IB of IRF3 dimerization (Native gel), p-IRF3, p-TBK1, p-IKKɛ, total IRF3, TBK1 or IKKɛ in HEK293T cells transfected with Flag-YAP2 expression plasmid and treated with SeV for the indicated time points. (d) IB of nuclear and cytoplasm fractions derived from HEK293T cells transfected with IRF3-Flag, IRF3-5D-Flag and YAP4-HA as indicated. (e) IB of TCL and immunoprecipitates derived from HEK293T cells transfected with Flag-Importin α5 or Flag-Importin β1 along with IRF3-5D-Myc. Data are representative of three independent experiments with similar results (b-f).
Supplementary Figure 5 YAP is promoted for lysosomal degradation by IKKɛ.
(a) Immunoblot of immunoprecipitates derived from HEK293T cells transfected with Flag-YAP2 plasmid and treated with SeV or poly (I:C) for 12 h. Cells were treated with NH4Cl (10 mM) for 4 h before harvest. (b) IB analysis for endogenous YAPs of control or IKKɛ depleted MCF10A cells treated with SeV for 12 h. (c) IB of HEK293T cells transfected with Flag-YAP4 and Myc-IKKɛ and treated with NH4Cl (10 mM), Chlq (100 μM), MG132 (10 μM) and control DMSO for 6 h. Data are representative of three independent experiments with similar results (a-c).
Supplementary Figure 6 YAP is phosphorylated by IKKɛ.
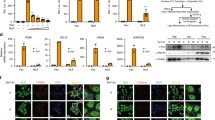
(a) Immunoblot of HEK293T cells transfected with expression vectors for Flag-YAP2 wt or Flag-YAP2 5SA (in which all the previously reported by Lats1 or 2 and CK1 phosphorylated Serines at positions 61, 109, 127, 128, 131, 163, 164, and 381 are changed to Alanine), with or without Myc-IKKɛ expression vectors as indicated. (b) Mass spectrometry identification of 4 phosphorylation sites in Flag-YAP2 5SA targeted by IKKɛ. (c) IB of HEK293T cells transfected with Myc-IKKɛ and Flag-YAP2 5SA derivatives carrying the indicated additional phosphorylation site mutations. (d) Sequence alignment of IKKɛ-mediated phosphorylation sites in YAP orthologues of different species. (e) IB of cell lysates of HEK293T cells transfected with Flag-YAP4 WT, 4SA or S403A mutant with or without Myc-IKKɛ expression plasmid as indicated. (f) Validation of the antibody against phosphor-Serine 403 of YAP. Parental, YAP1-deleted (KO) and YAP S403A-mutated (KI) HEK293T cells were treated with SeV for 8 h. Chlq (100 μM) were added to the cells 6 h before harvest. Cells were then harvested for immunoprecipitation with anti-YAP antibody followed by anti-p-S403 YAP immunoblot analysis. Data are representative of three independent experiments with similar results (a,c,e,f).
Supplementary Figure 7 IKKɛ-mediated phosphorylation of YAP at Ser403 is critical for the innate antiviral response.
(a) Immunoblot of IRF3 dimeration, and p-IRF3, p-TBK1, p-IKKɛ, total IRF3, TBK1 and IKKɛ in parental and YAP S403A knock-in (KI) A549 cells. Data are representative of three independent experiments with similar results. (b) qPCR analysis of IFNB1 mRNA levels (left panel) and ELISA analysis of IFN-β secretion in parental and YAP S403A-mutated A549 cells stimulated with SeV, VSV (MOI, 0.1) or transfected with cGAS and STING or poly (dA:dT) (right panel) for 12 h. qPCR and ELISA results are shown as mean ± sd of triplicates of at least two independent experiments.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7
Rights and permissions
About this article
Cite this article
Wang, S., Xie, F., Chu, F. et al. YAP antagonizes innate antiviral immunity and is targeted for lysosomal degradation through IKKɛ-mediated phosphorylation. Nat Immunol 18, 733–743 (2017). https://doi.org/10.1038/ni.3744
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.3744
This article is cited by
-
Overcoming BRAF and CDK4/6 inhibitor resistance by inhibiting MAP3K3-dependent protection against YAP lysosomal degradation
Experimental & Molecular Medicine (2024)
-
The role of non-canonical Hippo pathway in regulating immune homeostasis
European Journal of Medical Research (2023)
-
Tuning immunity through tissue mechanotransduction
Nature Reviews Immunology (2023)
-
MAVS deSUMOylation by SENP1 inhibits its aggregation and antagonizes IRF3 activation
Nature Structural & Molecular Biology (2023)
-
Non-hippo kinases: indispensable roles in YAP/TAZ signaling and implications in cancer therapy
Molecular Biology Reports (2023)