Abstract
The GTP-bound form of the Ran GTPase (RanGTP), produced around chromosomes, drives nuclear envelope and nuclear pore complex (NPC) re-assembly after mitosis. The nucleoporin MEL-28/ELYS binds chromatin in a RanGTP-regulated manner and acts to seed NPC assembly. Here we show that, upon mitotic NPC disassembly, MEL-28 dissociates from chromatin and re-localizes to spindle microtubules and kinetochores. MEL-28 directly binds microtubules in a RanGTP-regulated way via its C-terminal chromatin-binding domain. Using Xenopus egg extracts, we demonstrate that MEL-28 is essential for RanGTP-dependent microtubule nucleation and spindle assembly, independent of its function in NPC assembly. Specifically, MEL-28 interacts with the γ-tubulin ring complex and recruits it to microtubule nucleation sites. Our data identify MEL-28 as a RanGTP target that functions throughout the cell cycle. Its cell cycle-dependent binding to chromatin or microtubules discriminates MEL-28 functions in interphase and mitosis, and ensures that spindle assembly occurs only after NPC breakdown.
Similar content being viewed by others
Introduction
Chromosomes mediate the assembly of the nuclear envelope and NPCs in interphase and spindle assembly in mitosis1. RanGTP is produced around chromatin2 and plays a central role in both cell cycle stages. RanGTP binds to importin-α/β and dissociates nuclear localization signal (NLS)-containing proteins from importins3,4,5. In mitosis, liberated NLS proteins play distinct roles in spindle assembly around chromosomes6. To explore Ran-regulated spindle assembly factors, we previously purified and identified NLS-containing microtubule-associated proteins (MAPs) from Xenopus egg extracts7. These included the nucleoporin MEL-28 (maternal effect lethal-28)/ELYS (embryonic large molecule derived from yolk sac). MEL-28 is required for NPC assembly in human HeLa cells and Caenorhabditis elegans8,9,10. Further analysis using Xenopus egg extracts revealed that MEL-28 binds to chromatin in a RanGTP-dependent manner at early stages of NPC assembly11. MEL-28 interacts with the Nup107–160 complex (Nup for nucleoporin), an important building block of the NPC, and recruits the complex to chromatin. When NPCs disassemble in mitosis, MEL-28 partially localizes to kinetochores8,9,10. Its depletion from human cells or C. elegans embryos leads not only to nuclear pore defects in interphase but also to mitotic defects in chromosome condensation, kinetochore assembly, spindle assembly and chromosome segregation8,9,10. All these findings raise the question of whether MEL-28 affects mitosis directly or through its function in NPC assembly.
Here we find that MEL-28 dissociates from mitotic chromatin and re-localizes to the spindle as a MAP, and it is directly required for spindle assembly. MEL-28 promotes RanGTP-dependent γ-tubulin recruitment and microtubule nucleation. Our data show that MEL-28 is a RanGTP target that drives NPC assembly as cells exit mitosis and spindle assembly as cells enter mitosis.
Results
MEL-28 is a Ran-dependent MAP that relocates to the spindle
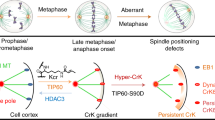
Our NLS-MAP purification7 contained MEL-28 with a high score, similar to known RanGTP-regulated MAPs such as TPX2 (Supplementary Table 1). Interestingly, Nup107 of the MEL-28-associated Nup107–160 complex (Nup160, Nup133, Nup107, Nup96, Nup85, Nup43, Nup37, Sec13 and Seh1, see Supplementary Table 1) was identified with a lower score, suggesting lower abundance. We examined the ability of MEL-28, included in HeLa nuclear extracts, to bind microtubules in a sedimentation assay. After sedimentation of taxol-stabilized microtubules, human MEL-28 was efficiently depleted from the extract and recovered in the pellet, similar to the nuclear MAP ISWI/SNF2H (Fig. 1a)7. In contrast, Nup107–160 components were not depleted (Fig. 1a) and were only detectable in the pellet when a 20-fold excess was probed (Supplementary Fig. 1a). MEL-28 was recovered in the pellet only in the presence of microtubules and under physiological salt concentration (Fig. 1b). Microtubule binding of MEL-28 was inhibited by recombinant importin-α and -β, but it was restored by further addition of RanGTP, similar to microtubule binding of ISWI (positive control) but not of ch-TOG (negative control) (Supplementary Fig. 1b).
(a) HeLa nuclear extract (NE) was incubated with taxol-stabilized microtubules (MTs) and sedimented. Supernatant (s) and pellet (p) fractions were analysed by immunoblotting. (b) NE was incubated with or without MTs and with or without additional 0.5 M NaCl. After centrifugation, pellets were analysed by immunoblotting or Ponceau staining. (c) GST-fused Xenopus MEL-28 aa 1,441–2,201 was incubated with MTs in the presence or absence of importin-α, importin-β and RanGTP. After centrifugation, samples were analysed by Coomassie staining. (d) GFP, GFP-fused aa 1,441–2,201 or histidine-tagged ISWI were incubated with or without Cy3-labelled, taxol-stabilized MTs at RT for 10 min. Note that his-ISWI is a negative control that bundles MTs but does not show green signals. Bar, 20 μm. (e,f) Sperm nuclei in Xenopus egg extract were cycled to interphase and then back to mitosis. (e) At indicated time points, chromatin was isolated and immunoblotted for MEL-28, ISWI and RCC1. Coomassie staining of the gel shows histones as an indicator of chromatin recovery. (f) Interphase nuclei and mitotic spindles were stained with a Xenopus MEL-28 antibody (red) and Alexa488-labelled tubulin (green), as described in the methods. DNA was stained with Hoechst (blue). Bar, 20 μm. (g) CSF extract was incubated with RanGTP and Alexa488-tubulin (green). Ran-induced asters and spindles were stained with an MEL-28 antibody (red). Bars, 20 μm. (h) HeLa cells were pre-extracted with Triton X-100, fixed and stained with human MEL-28 antibodies against aa 1,572–2,266 (ref. 11) or aa 1,208–1,800 fragments (green). Cells were also stained for α-tubulin (red) and DNA (blue). Bars, 10 μm. (i) Human MEL-28 antibody was generated in rabbits against recombinant aa 1,208–1,800. HeLa cell lysate was blotted with the antibody showing a monospecific signal at ~250 kD.
ISWI binds to microtubules via the region containing chromatin-binding domains and an NLS7, which prompted us to test a potential direct microtubule-binding site of MEL-28 in its C terminus containing an AT-hook (AT-rich DNA-binding motif) and an NLS (Supplementary Fig. 1c)12. A MEL-28 C-terminal fragment comprising aa 1,441–2,201 bound to microtubules. Binding was inhibited by importin-α/β and restored by RanGTP (Fig. 1c). GFP-fused aa 1,441–2,201 bound along microtubules and bundled them (Fig. 1d). Strikingly, aa 1,602–2,120, lacking the AT-hook and NLS, did not bind to microtubules, whereas aa 1,993–2,201 bound (Supplementary Fig. 1d). Therefore, the C-terminal region containing AT-hook and NLS is the RanGTP-dependent microtubule-binding site of MEL-28.
It has been reported previously that only a subpopulation of MEL-28 interacts with the Nup107–160 complex11. Consistently, when we immunodepleted MEL-28 from HeLa nuclear extracts, the Nup107–160 components were only partially depleted (Supplementary Fig. 1e). Importantly, microtubule binding of the remaining Nup107–160 components was inhibited in the sedimentation assay (Supplementary Fig. 1e), indicating that Nup107–160 complex binding to microtubules occurs via MEL-28.
MEL-28 binds to microtubules via its chromatin-binding region, raising the question of how the distribution of MEL-28 between chromatin and microtubules is regulated during the cell cycle. By analysing chromatin re-isolated from cycling Xenopus egg extracts, we found that MEL-28 completely dissociates from chromatin on mitotic entry (Fig. 1e)13. In interphase egg extract, MEL-28 antibodies stained the rim and interior of in vitro-assembled nuclei (Fig. 1f)8,11. In the mitotic extract, we detected MEL-28 on kinetochores and spindle microtubules, with enrichment on spindle poles (Fig. 1f). Microtubule and pole localization was also observed in RanGTP-induced asters or spindle-like structures assembled in M-phase-arrested egg extracts in the absence of chromatin and kinetochores (Fig. 1g). MEL-28 has been shown to localize to kinetochores in mitotic cells8,11. We re-examined the localization of MEL-28 in HeLa cells. When soluble MEL-28 was extracted before fixation, residual MEL-28 was clearly detected around spindle poles, as well as on kinetochores (Fig. 1h,i). Therefore, spindle pole and kinetochore localization of MEL-28 is conserved in Xenopus and human cells.
Independent MEL-28 functions in spindle and NPC assembly
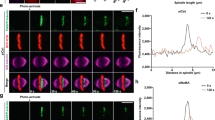
We next immunodepleted MEL-28 from M-phase-arrested Xenopus egg extracts and tested spindle formation around sperm nuclei (Fig. 2a)14. Nup107–160 components, as expected, were only partially depleted. 10 min after incubation with sperm, MEL-28-depleted extracts did assemble microtubules but with significantly lower effectiveness than controls (Fig 2b,c). After 80 min, control extracts assembled bipolar spindles on sperm chromatin with ~50% frequency. In depleted extracts, the vast majority of sperm had no microtubules (Fig. 2b–d), indicating that initially generated microtubules were unstable. Importantly, when we expressed MEL-28 in depleted extracts using a corresponding mRNA, spindle assembly was fully restored yielding spindles with normal bipolarity and microtubule density (Fig. 2e–g).
(a) CSF extract was immunodepleted using rabbit IgG (mock) or a Xenopus MEL-28 antibody, and analysed by immunoblotting. α-tubulin as a control protein was not depleted. Numbers (%, ±s.d. from three independent experiments) represent protein levels after MEL-28 depletion, quantified with non-saturated blots. (b–d) Spindle assembly around sperm nuclei was examined in control or MEL-28-depleted CSF extracts. At the indicated time point, aliquots were fixed and spun down on coverslips. (b) Representative images: blue, DNA; red, Cy3-labelled tubulin. (c) The microtubule (MT) intensity around sperm was quantified. We defined the MT intensity in control extracts as 1. n>20 structures. Error bars, s.d. (d) Quantification of spindle phenotypes over the total number of sperm counted. n>50 sperm. Error bars, s.d. from three experiments. (e–g) Control or MEL-28-depleted CSF extracts were incubated with or without mRNA encoding MEL-28 at 20 °C for 60 min, and spindle assembly was assayed as in b. (e) MEL-28 depletion and expression was analysed by immunoblotting. (f) Representative images. (g) Quantification of the MT intensity around sperm. Box-and-whisker plots show median (horizontal line), interquartile range (box) and maximum/minimum range (whiskers). n>30 structures from three independent experiments. ***P<0.001, ****P<0.0001 (Student’s t-test, two-tailed). Bars, 20 μm.
MEL-28 is needed for chromatin-driven microtubule nucleation
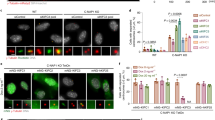
MEL-28 is concentrated on kinetochores and might thus primarily function there. To examine this, we assembled spindles around chromatin-coated beads that lack both kinetochores and centrosomes15. On MEL-28 depletion, microtubule assembly around chromatin beads was severely inhibited (Fig. 3a–c), as for sperm nuclei. No microtubules were nucleated around chromatin beads after 10 min (Fig. 3a,b). RanGTP-induced aster and spindle formation16 was also abolished in the absence of MEL-28 (Fig. 3d,e). In contrast, the addition of DMSO to both mock and MEL-28-depleted extracts induced similar numbers of aster-like structures (Supplementary Fig. 2).
(a–c) Chromatin bead spindles were assembled in mock- or MEL-28-depleted CSF extracts. At the indicated time points, aliquots were fixed and spun down on coverslips. (a) Representative images: red, microtubules (MTs); blue, DNA. (b) Quantification of the MT intensity around DNA beads. n>20 DNA bead clusters containing 15–40 beads, which frequently formed bipolar spindles in control extracts. Error bars, s.d. (c) Quantification of spindle phenotypes. n>50 DNA bead clusters. Error bars, s.d. from two experiments. (d,e) RanGTP-mediated MT assembly was examined in control or MEL-28-depleted extracts. Samples were squash-fixed at the indicated time points. (d) Representative MT images. (e) Numbers of MT structures and spindle-like structures were counted in 100 randomly selected fields with a 63 × objective. Error bars, s.d. from four experiments. (f,g) Centrosome-mediated MT assembly was examined in control or MEL-28-depleted extracts. Samples were fixed and spun down on coverslips. Note that this assay was performed in TPX2-depleted extracts that do not nucleate self-organized MTs and allow MT nucleation exclusively from centrosomes. (f) Representative images of asters. (g) MT length and intensity in centrosomal asters were quantified31. n>20 asters. Error bars, s.d. Not significant (NS), P>0.05. *P<0.05, **P<0.01, ***P<0.001, ****P<0.0001 (Student’s t-test, two-tailed). Scale bars, 20 μm.
We next analysed whether MEL-28 also regulates microtubule stability. Incubation of centrosomes in control extracts nucleated microtubule asters, and the addition of RanGTP increased microtubule length and density17 (Fig. 3f,g). In MEL-28-depleted extracts, microtubule length increased on RanGTP addition as in the control. In contrast, microtubule intensity was less effectively stimulated by RanGTP (Fig. 3f,g). Therefore, MEL-28 is required for RanGTP-dependent microtubule nucleation, but not stabilization.
MEL-28 is critical in Ran-dependent recruitment of γ-tubulin
RanGTP-dependent microtubule nucleation requires the γ-tubulin ring complex (γTuRC)18. Immunoprecipitation of MEL-28 identified γTuRC and Nup107–160 components as main interaction partners (Fig. 4a; mass spectrometry in Supplementary Table 2). The interaction between MEL-28 and γTuRC was confirmed by γ-tubulin immunoprecipitation (Fig. 4a). In contrast, we could not detect interaction between MEL-28 and HURP or TPX2 (ref. 19; Fig. 4a).
(a) MEL-28 interacts with γTuRC. CSF extract was incubated with protein A beads covalently coupled to control IgG or antibodies against MEL-28, HURP, TPX2 or γ-tubulin. The beads were washed and immunoprecipitates (IP) were analysed by immunoblotting. (b) Centrosomal asters were assembled as in Fig. 3f, but subsequently immunostained for γ-tubulin (green). Red, microtubules (MTs). Insets show γ-tubulin signal on centrosomes with the same magnification. Scale bar, 20 μm. γ-Tubulin levels at centrosomes were quantified. n>20 asters. Error bars, s.d. (c) Mock or MEL-28-depleted CSF extracts were incubated with sperm and Cy3-tubulin (red) at 20 °C for 10 min. The samples were fixed, spun down on coverslips and stained for γ-tubulin (green). Scale bar, 20 μm. γ-Tubulin levels at centrosomes were quantified in a 3-μm-diameter circle. n>20 asters assembled around sperm. Error bars, s.d. ****P<0.0001 (Student’s t-test, two-tailed). (d) Model to highlight the mutually exclusive functions of MEL-28 to assemble NPCs in interphase or to promote MT nucleation in mitosis (details see text). (e) Domain structure of MEL-28 orthologues. The tree shows the phylogenetic distribution of the selected species according to NCBI taxonomy. Thick black lines indicate the length of each protein. Schematic representation displays β-propeller (blue), coiled-coil domain (yellow), MEL-28/ELYS domain (red) and AT-hook (green). Species known to disassemble NPCs are displayed in orange, and those that keep NPCs assembled during mitosis are in purple (black: no information).
We therefore examined the levels of γ-tubulin on centrosomal asters. RanGTP stimulated γ-tubulin recruitment to centrosomes in control extracts17, but not in MEL-28-depleted extracts (Fig. 4b). Moreover, γ-tubulin, essential for microtubule nucleation, was recruited to sperm centrosomes at early stages of mitosis in control extracts (Fig. 4c)18. MEL-28 depletion abolished this recruitment (Fig. 4c). These results clearly indicate that MEL-28 binds to microtubules in a RanGTP-dependent manner and recruits γ-tubulin for further microtubule nucleation.
Discussion
Experiments in Xenopus egg extracts showed that at early stages of NPC assembly MEL-28 binds to chromatin in a RanGTP-dependent manner11. Chromatin binding is via the C-terminal region containing an AT-hook and an NLS, and is required for the recruitment of other nucleoporins and progression of NPC assembly20 (Fig. 4d). Here we find that on mitotic entry and NPC disassembly, MEL-28 dissociates from chromatin and binds microtubules via the same C-terminal region in a RanGTP-dependent manner (Fig. 4d). This allows MEL-28 to play its roles in microtubule assembly and spindle formation. MEL-28 binding to chromatin or microtubules using the same C-terminal region may discriminate its function in interphase and mitosis in a mutually exclusive way. Interestingly, yeast cells undergo mitosis without disassembling NPCs and express ELY5 (35 kD) as an orthologue of metazoan MEL-28 (>200 kD)21. ELY5 comprises only the central α-helical MEL-28/ELYS domain and lacks all other domains (β-propeller, coiled-coil and unstructured C-terminal domain containing the AT-hook and the NLS) (Fig. 4e and Supplementary Fig. 3). Yeast ELY5 is a component of NPCs, but, in contrast to metazoans, is not required for NPC assembly21. Fungi with the minimal MEL-28 domain also do not disassemble NPCs during mitosis22 (Fig. 4e). Taken together, these data suggest that the RanGTP-dependent functions of MEL-28 in interphase and mitosis evolved with NPC disassembly and open mitosis.
MEL-28 is specifically required for RanGTP-dependent microtubule nucleation in egg extract, like TPX2 and HURP19. MEL-28, however, does not associate with these two proteins, but interacts with γTuRC and the Nup107–160 complex. MEL-28, TPX2 and HURP may be involved in different steps of RanGTP-dependent microtubule nucleation23. TPX2 depletion induces the most severe defects in Ran-induced microtubule nucleation3, whereas MEL-28 depletion results in the most severe defects in sperm spindle assembly. We hypothesize that TPX2 is critical for de novo formation of microtubules in the vicinity of chromatin. MEL-28 binds to pre-existing microtubules, nucleated by TPX2 or from centrosomes, in a RanGTP-dependent manner and recruits γ-tubulin to promote additional microtubule nucleation. MEL-28 thus drives positive feedback in RanGTP-mediated microtubule nucleation24,25. The nearly complete absence of microtubules around chromatin in MEL-28-depleted extracts may be attributable to the chromatin-bound protein Dppa2, which locally inhibits microtubule assembly26. Dppa2 on chromatin may effectively depolymerize microtubules nucleated by TPX2 and centrosomes in the absence of MEL-28. In contrast, it hardly affects further amplified microtubules in the presence of MEL-28.
γTuRC primarily localizes to centrosomes and spindle microtubules under physiological conditions18 and accumulates at kinetochores only when microtubule attachments are disrupted27. Our data support the idea that MEL-28 primarily functions in microtubule nucleation at the spindle. It has been shown previously that the Nup107–160 complex partially localizes to the spindle and is required for spindle assembly in egg extracts28. We provide strong evidence that MEL-28 is the MAP recruiting the Nup107–160 complex and γTuRC to microtubules, and is the key factor for RanGTP-dependent microtubule nucleation.
In summary, we have discovered a new role for MEL-28 as a RanGTP-dependent MAP required for microtubule nucleation and spindle assembly. Our data highlight that the cell cycle-dependent association of MEL-28 exclusively drives NPC assembly at the end of mitosis and spindle assembly in early mitosis (Fig. 4d).
Methods
Xenopus egg extracts
Cytostatic factor-arrested M-phase Xenopus laevis egg extracts (CSF extracts) were prepared29,30. Xenopus eggs were dejellied by cysteine treatment, washed with XB buffer (10 mM K–HEPES, 100 mM KCl, 1 mM MgCl2, 0.1 mM CaCl2 and 50 mM sucrose, pH 7.7) and subsequently with CSF–XB buffer (10 mM K–HEPES, 100 mM KCl, 3 mM MgCl2, 0.1 mM CaCl2, 50 mM sucrose and 5 mM EGTA, pH 7.7) and crushed by centrifugation at 20,000 g for 20 min at 16 °C. The straw-coloured middle layer was recovered as a CSF extract. Endogenous MEL-28 was depleted from CSF extracts by two rounds of incubation with 60% (vol/vol) Dynabeads Protein A (Life Technologies) coupled with Xenopus MEL-28 antibody11 (0.3 μg antibody applied per 1 μl beads). CSF extracts were supplemented with Cy3-labelled tubulin and incubated with sperm nuclei or chromatin beads for spindle assembly30. Microtubule density around sperm or beads was quantified using Matlab (The MathWorks)31. For rescue experiments, mRNA encoding Xenopus MEL-28 (GenBank accession KJ026765) was prepared using the mMESSAGE mMachine kit (Life Technologies) and added to extracts at a concentration of 300 ng μl−1.
Recombinant proteins and antibodies
For recombinant fragments of Xenopus MEL-28 (KJ026765), cDNAs encoding aa 1,441–2,201 and aa 1,993–2,201 were subcloned into pGEX-6P-1. The GST fusion proteins were expressed in E. coli and purified with Glutathione Sepharose (GE Healthcare Life Sciences). His-tagged aa 1,602–2,120 was expressed in E. coli and purified with Ni-NTA (Qiagen)11. His, GFP-tagged aa 1,441–2,201 was constructed in pET28a (Novagen), expressed in E. coli and purified with Ni-NTA and Superose 6 (GE Healthcare Life Sciences). His-importin-α, His-importin-β and His-RanQ69L-GTP were expressed in E. coli and purified with TALON beads (Clontech)31.
The following published and commercial antibodies were used: Xenopus MEL-28 antibody, human MEL-28 antibody against aa 1,572–2,266, Xenopus 160 antibody11, human Nup107, Nup133 and Nup160 antibodies (Abcam ab85916, ab57645 and ab74147, respectively), γ-tubulin antibody from mice (Sigma T6557), γ-tubulin antibody from rabbits32, Xgrip109 and 210 antibodies33. The human MEL-28 antibody was produced in rabbits against recombinant aa 1,208–1,800, Xenopus Nup107 antibody was produced against aa 1–164 and Xenopus Nup133 antibody was produced against aa 640–1,060.
Microtubule assays in vitro and in egg extracts
Microtubule sedimentation assay: HeLa nuclear extract (4C Biotech) was diluted with CSF–XB buffer to 1 mg ml−1 and centrifuged at 100,000 g for 10 min at 20 °C. The supernatant was incubated with 2 μM taxol-stabilized microtubules at room temperature (RT) for 15 min in the presence of 1 mM GTP and 10 μM taxol. The samples were centrifuged at 100,000 g for 10 min at 20 °C, and the supernatant and pellet were analysed by SDS–PAGE and immunoblotting. Recombinant MEL-28 fragments were incubated with MTs in the same way but in the presence of 0.05 mg ml−1 BSA.
Microscopy-based microtubule-binding assay was performed in vitro7. GFP MEL-28 aa 1,441–2,201 (1 μM) was incubated with Cy3-labelled, taxol-stabilized MTs (0.3 μM) at RT for 10 min in BRB80 buffer (80 mM K-PIPES, 1 mM MgCl2, 1 mM EGTA, pH6.8) supplemented with 20 μM taxol. The samples were squashed with fixative.
The RanGTP-dependent microtubule nucleation assay16: CSF extracts were incubated with 15 μM RanQ69L-GTP and 1 μM Cy3-tubulin at 20 °C for 80 min. The samples (Ran spindles and asters) were squashed with fixative.
The RanGTP-dependent microtubule stabilization assay31: TPX2-depleted CSF extracts were incubated with 1 μM Cy3-tubulin and 2,000 centrosomes per μl in the presence or absence of 15 μM RanQ69L-GTP at 20 °C for 30 min. The samples (centrosomal asters) were fixed and spun down on coverslips.
Immunofluorescence
CSF extract was supplemented with sperm nuclei and Alexa 488-labelled tubulin, and driven into interphase by adding calcium and incubating at 20 °C for 90 min. Samples were cycled into mitosis by adding a fresh CSF extract and incubating at 20 °C for 80 min. Cy3-labelled Xenopus MEL-28 antibody was added to the extract at a 5 ng μl−1 concentration 10 min before fixation. Interphase nuclei and mitotic spindles were fixed and spun down on coverslips. DNA was stained with Hoechst 33342.
HeLa cells were first permeabilized with 0.1% Triton X-100 in PHEM buffer (60 mM PIPES, 20 mM HEPES, pH 6.9, 10 mM EGTA, 4 mM MgSO4) at RT for 5 min and then fixed with −20 °C methanol for 5 min28. The cells were stained with human MEL-28 antibody (1 μg ml−1, against aa 1,572–2,266 or aa 1,208–1,800) and anti-α-tubulin DM1A (1:1,000; Sigma T9026) followed by Alexa 488-labelled anti-rabbit IgG and Alexa 568-labelled anti-mouse IgG (Life Technologies). DNA was stained with Hoechst.
Chromatin isolation
CSF extract supplemented with sperm and Cy3-tubulin were driven into interphase and cycled into mitosis as described above. At each time point, an aliquot was taken and chromatin was isolated34. Samples were diluted with 5 volumes of XB buffer and centrifuged at 5,000 g for 12 min through a 0.7-M sucrose cushion. After removing the supernatant, nuclear pellets were resuspended in XB, 0.3% Triton X-100 and incubated for 5 min on ice. After an additional 10,000g centrifugation for 2 min, the chromatin fraction (pellet) was recovered.
Microscopy
Fluorescence images were acquired using a Zeiss Cell Observer microscope, a Plan-Apo 63 × NA 1.4 oil objective, an AxioCam MRm camera and the AxioVision software. Images were also acquired at 0.5-μm Z steps using a Leica SP5 confocal microscope equipped with a 63 × Plan-apochromat NA 1.4 oil-immersion objective. Maximum intensity projections were obtained using ImageJ (NIH).
Immunoprecipitations
Dynabeads Protein A were coupled with various purified antibodies and cross-linked with dimethyl pimelimidate (Sigma)35. Each bead sample (60 μl slurry) was incubated with CSF extracts (100 μl) at 4 °C for 60 min and washed twice with CSF–XB and twice with CSF–XB containing 0.5 M KCl and 0.1% Triton X-100. The immunoprecipitates were separated by SDS–PAGE for immunoblotting. We also analysed control and MEL-28 immunoprecipitates by mass spectrometry32. In brief, proteins of the immunoprecipitates were separated by SDS–PAGE, and proteins in-gel were digested by trypsin. Samples were analysed using an ESI LTQ Orbitrap mass spectrometer (Thermo Fisher, Dreieich, Germany).
Bioinformatics
Homologues of X. laevis MEL-28 were identified with a PSI-BLAST three-iteration search against the non-redundant protein sequences database, performed on the NCBI server ( http://blast.ncbi.nlm.nih.gov/Blast.cgi) with default parameters. Representative protein sequences were then selected and aligned using T-Coffee with default parameters36. The resulting alignment was visualized with Jalview37 coloured with Clustal X colour scheme.
A hidden Markov model was built from the annotated AT-hook sequences using the HHsuite 2.0 package38, which was used to identify this DNA-binding domain in other homologous proteins with HHsearch. Secondary structures were predicted by PSIPRED39, whereas disorder was predicted with IUPred40. The phylogenetic tree was generated with iTOL41 based on the eukaryotic NCBI taxonomy tree.
Western blots
Original scans of the cropped images in the main figures (Figs 1a,e, 2a and 4a) are presented in Supplementary Fig. 4.
Additional information
How to cite this article: Yokoyama, H. et al. The nucleoporin MEL-28 promotes RanGTP-dependent γ-tubulin recruitment and microtubule nucleation in mitotic spindle formation. Nat. Commun. 5:3270 doi: 10.1038/ncomms4270 (2014).
References
Guttinger, S., Laurell, E. & Kutay, U. Orchestrating nuclear envelope disassembly and reassembly during mitosis. Nat. Rev. Mol. Cell Biol. 10, 178–191 (2009).
Kalab, P., Weis, K. & Heald, R. Visualization of a Ran-GTP gradient in interphase and mitotic Xenopus egg extracts. Science 295, 2452–2456 (2002).
Gruss, O. J. et al. Ran induces spindle assembly by reversing the inhibitory effect of importin on TPX2 activity. Cell 104, 83–93 (2001).
Nachury, M. V. et al. Importin beta is a mitotic target of the small GTPase Ran in spindle assembly. Cell 104, 95–106 (2001).
Wiese, C. et al. Role of importin-beta in coupling Ran to downstream targets in microtubule assembly. Science 291, 653–656 (2001).
Meunier, S. & Vernos, I. Microtubule assembly during mitosis - from distinct origins to distinct functions? J. Cell Sci. 125, 2805–2814 (2012).
Yokoyama, H., Rybina, S., Santarella-Mellwig, R., Mattaj, I. W. & Karsenti, E. ISWI is a RanGTP-dependent MAP required for chromosome segregation. J. Cell Biol. 187, 813–829 (2009).
Rasala, B. A., Orjalo, A. V., Shen, Z., Briggs, S. & Forbes, D. J. ELYS is a dual nucleoporin/kinetochore protein required for nuclear pore assembly and proper cell division. Proc. Natl Acad. Sci. USA 103, 17801–17806 (2006).
Galy, V., Askjaer, P., Franz, C., Lopez-Iglesias, C. & Mattaj, I. W. MEL-28, a novel nuclear-envelope and kinetochore protein essential for zygotic nuclear-envelope assembly in C. elegans. Curr. Biol. 16, 1748–1756 (2006).
Fernandez, A. G. & Piano, F. MEL-28 is downstream of the Ran cycle and is required for nuclear-envelope function and chromatin maintenance. Curr. Biol. 16, 1757–1763 (2006).
Franz, C. et al. MEL-28/ELYS is required for the recruitment of nucleoporins to chromatin and postmitotic nuclear pore complex assembly. EMBO Rep. 8, 165–172 (2007).
Lau, C. K. et al. Transportin regulates major mitotic assembly events: from spindle to nuclear pore assembly. Mol. Biol. Cell 20, 4043–4058 (2009).
Gillespie, P. J., Khoudoli, G. A., Stewart, G., Swedlow, J. R. & Blow, J. J. ELYS/MEL-28 chromatin association coordinates nuclear pore complex assembly and replication licensing. Curr. Biol. 17, 1657–1662 (2007).
Sawin, K. E. & Mitchison, T. J. Mitotic spindle assembly by two different pathways in vitro. J. Cell Biol. 112, 925–940 (1991).
Heald, R. et al. Self organization of microtubules into bipolar spindles around artificial chromosomes in Xenopus egg extracts. Nature 382, 420–425 (1996).
Carazo-Salas, R. E. et al. Generation of GTP-bound Ran by RCC1 is required for chromatin-induced mitotic spindle formation. Nature 400, 178–181 (1999).
Carazo-Salas, R. E., Gruss, O. J., Mattaj, I. W. & Karsenti, E. Ran-GTP coordinates regulation of microtubule nucleation and dynamics during mitotic-spindle assembly. Nat. Cell Biol. 3, 228–234 (2001).
Wilde, A. & Zheng, Y. Stimulation of microtubule aster formation and spindle assembly by the small GTPase Ran. Science 284, 1359–1362 (1999).
Casanova, C. M., Rybina, S., Yokoyama, H., Karsenti, E. & Mattaj, I. W. Hepatoma up-regulated protein is required for chromatin-induced microtubule assembly independently of TPX2. Mol. Biol. Cell 19, 4900–4908 (2008).
Rasala, B. A., Ramos, C., Harel, A. & Forbes, D. J. Capture of AT-rich chromatin by ELYS recruits POM121 and NDC1 to initiate nuclear pore assembly. Mol. Biol. Cell 19, 3982–3996 (2008).
Bilokapic, S. & Schwartz, T. U. Molecular basis for Nup37 and ELY5/ELYS recruitment to the nuclear pore complex. Proc. Natl Acad. Sci. USA 109, 15241–15246 (2012).
De Souza, C. P., Osmani, A. H., Hashmi, S. B. & Osmani, S. A. Partial nuclear pore complex disassembly during closed mitosis in Aspergillus nidulans. Curr. Biol. 14, 1973–1984 (2004).
Teixido-Travesa, N., Roig, J. & Luders, J. The where, when and how of microtubule nucleation - one ring to rule them all. J. Cell Sci. 125, 4445–4456 (2012).
Clausen, T. & Ribbeck, K. Self-organization of anastral spindles by synergy of dynamic instability, autocatalytic microtubule production, and a spatial signaling gradient. PLoS One 2, e244 (2007).
O'Connell, C. B. & Khodjakov, A. L. Cooperative mechanisms of mitotic spindle formation. J. Cell Sci. 120, 1717–1722 (2007).
Xue, J. Z., Woo, E. M., Postow, L., Chait, B. T. & Funabiki, H. Chromatin-bound Xenopus dppa2 shapes the nucleus by locally inhibiting microtubule assembly. Dev. Cell 27, 47–59 (2013).
Mishra, R. K., Chakraborty, P., Arnaoutov, A., Fontoura, B. M. & Dasso, M. The Nup107-160 complex and gamma-TuRC regulate microtubule polymerization at kinetochores. Nat. Cell Biol. 12, 164–169 (2010).
Orjalo, A. V. et al. The Nup107-160 nucleoporin complex is required for correct bipolar spindle assembly. Mol. Biol. Cell 17, 3806–3818 (2006).
Murray, A. Cell cycle extracts. inXenopus laevis: Practical Uses in Cell and Molecular Biology Vol. 36, eds Kay B. K., Peng H. B. 581–605Academic press, inc., San Diego, New york, Boston, London, Sydney, Tokyo, Toronto (1991).
Hannak, E. & Heald, R. Investigating mitotic spindle assembly and function in vitro using Xenopus laevis egg extracts. Nat. Protoc. 1, 2305–2314 (2006).
Yokoyama, H. et al. Cdk11 is a RanGTP-dependent microtubule stabilization factor that regulates spindle assembly rate. J. Cell Biol. 180, 867–875 (2008).
Barenz, F. et al. The centriolar satellite protein SSX2IP promotes centrosome maturation. J. Cell Biol. 202, 81–95 (2013).
Zhang, L., Keating, T. J., Wilde, A., Borisy, G. G. & Zheng, Y. The role of Xgrip210 in gamma-tubulin ring complex assembly and centrosome recruitment. J. Cell Biol. 151, 1525–1536 (2000).
Demeret, C., Bocquet, S., Lemaitre, J. M., Francon, P. & Mechali, M. Expression of ISWI and its binding to chromatin during the cell cycle and early development. J. Struct. Biol. 140, 57–66 (2002).
Harlow, E. & Lane, D. Using Antibodies a Laboratory Manual Cold Spring Harbor Laboratory, Cold Spring Harbor, New York (1999).
Notredame, C., Higgins, D. G. & Heringa, J. T-Coffee: a novel method for fast and accurate multiple sequence alignment. J. Mol. Biol. 302, 205–217 (2000).
Waterhouse, A. M., Procter, J. B., Martin, D. M., Clamp, M. & Barton, G. J. Jalview Version 2--a multiple sequence alignment editor and analysis workbench. Bioinformatics 25, 1189–1191 (2009).
Soding, J. Protein homology detection by HMM-HMM comparison. Bioinformatics 21, 951–960 (2005).
Jones, D. T. Protein secondary structure prediction based on position-specific scoring matrices. J. Mol. Biol. 292, 195–202 (1999).
Dosztanyi, Z., Csizmok, V., Tompa, P. & Simon, I. IUPred: web server for the prediction of intrinsically unstructured regions of proteins based on estimated energy content. Bioinformatics 21, 3433–3434 (2005).
Letunic, I. & Bork, P. Interactive Tree Of Life (iTOL): an online tool for phylogenetic tree display and annotation. Bioinformatics 23, 127–128 (2007).
Acknowledgements
We thank Y. Zheng for antibodies against XGrips and K. Crnokic for maintaining the frogs. We also thank the Advanced Light Microscopy and Proteomics Core Facilities at EMBL and the Light Microscopy and Mass Spectrometry Facilities at ZMBH. This study was supported by DFG (1737/4-3) and a Start-Professorship of the German Excellence Initiative (ZUK49/1 TP 1-16) to O.J.G.
Author information
Authors and Affiliations
Contributions
H.Y. designed the study. H.Y., I.W.M. and O.J.G. supervised the research. H.Y., B.K., R.W., F.C.D. and O.J.G. performed the experiments. B.K. and R.W. prepared recombinant proteins and antibodies. J.C.G.-S. and D.P.D. conducted the bioinformatics analyses. H.Y. and O.J.G. wrote the paper and I.W.M. revised it. H.Y. and O.J.G. jointly managed the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures 1-4, Supplementary Tables 1-2 and Supplementary References (PDF 623 kb)
Rights and permissions
About this article
Cite this article
Yokoyama, H., Koch, B., Walczak, R. et al. The nucleoporin MEL-28 promotes RanGTP-dependent γ-tubulin recruitment and microtubule nucleation in mitotic spindle formation. Nat Commun 5, 3270 (2014). https://doi.org/10.1038/ncomms4270
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/ncomms4270
This article is cited by
-
Mechanisms underlying spindle assembly and robustness
Nature Reviews Molecular Cell Biology (2023)
-
The γ-tubulin meshwork assists in the recruitment of PCNA to chromatin in mammalian cells
Communications Biology (2021)
-
Multidomain ribosomal protein trees and the planctobacterial origin of neomura (eukaryotes, archaebacteria)
Protoplasma (2020)
-
Two-step interphase microtubule disassembly aids spindle morphogenesis
BMC Biology (2018)
-
γ-tubulin as a signal-transducing molecule and meshwork with therapeutic potential
Signal Transduction and Targeted Therapy (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.