Abstract
Mitogen-activated protein kinase (MAPK) cascades play central roles in innate immune signalling networks in plants and animals1,2. In plants, however, the molecular mechanisms of how signal perception is transduced to MAPK activation remain elusive1. Here we report that pathogen-secreted proteases activate a previously unknown signalling pathway in Arabidopsis thaliana involving the Gα, Gβ, and Gγ subunits of heterotrimeric G-protein complexes, which function upstream of an MAPK cascade. In this pathway, receptor for activated C kinase 1 (RACK1) functions as a novel scaffold that binds to the Gβ subunit as well as to all three tiers of the MAPK cascade, thereby linking upstream G-protein signalling to downstream activation of an MAPK cascade. The protease–G-protein–RACK1–MAPK cascade modules identified in these studies are distinct from previously described plant immune signalling pathways such as that elicited by bacterial flagellin, in which G proteins function downstream of or in parallel to an MAPK cascade without the involvement of the RACK1 scaffolding protein. The discovery of the new protease-mediated immune signalling pathway described here was facilitated by the use of the broad host range, opportunistic bacterial pathogen Pseudomonas aeruginosa. The ability of P. aeruginosa to infect both plants and animals makes it an excellent model to identify novel immunoregulatory strategies that account for its niche adaptation to diverse host tissues and immune systems.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Tena, G., Boudsocq, M. & Sheen, J. Protein kinase signaling networks in plant innate immunity. Curr. Opin. Plant Biol. 14, 519–529 (2011)
Arthur, J. S. & Ley, S. C. Mitogen-activated protein kinases in innate immunity. Nature Rev. Immunol. 13, 679–692 (2013)
Millet, Y. A. et al. Innate immune responses activated in Arabidopsis roots by microbe-associated molecular patterns. Plant Cell 22, 973–990 (2010)
Denoux, C. et al. Activation of defense response pathways by OGs and flg22 elicitors in Arabidopsis seedlings. Mol. Plant 1, 423–445 (2008)
Traidej, M., Marquart, M. E., Caballero, A. R., Thibodeaux, B. A. & O’Callaghan, R. J. Identification of the active site residues of Pseudomonas aeruginosa protease IV. J. Biol. Chem. 278, 2549–2553 (2003)
Urano, D., Chen, J.-G., Botella, J. R. & Jones, A. M. Heterotrimeric G protein signalling in the plant kingdom. Open Biol. 3, 120186 (2013)
Ullah, H. et al. Structure of a signal transduction regulator, RACK1, from Arabidopsis thaliana. Protein Sci. 17, 1771–1780 (2008)
Dell, E. J. et al. The βγ subunit of heterotrimeric G proteins interacts with RACK1 and two other WD repeat proteins. J. Biol. Chem. 277, 49888–49895 (2002)
Nakashima, A. et al. RACK1 functions in rice innate immunity by interacting with the Rac1 immune complex. Plant Cell 20, 2265–2279 (2008)
Chen, J.-G. et al. RACK1 mediates multiple hormone responsiveness and developmental processes in Arabidopsis. J. Exp. Bot. 57, 2697–2708 (2006)
Asai, T. et al. MAP kinase signalling cascade in Arabidopsis innate immunity. Nature 415, 977–983 (2002)
Suarez-Rodriguez, M. C. et al. MEKK1 is required for flg22-induced MPK4 activation in Arabidopsis plants. Plant Physiol. 143, 661–669 (2007)
MAPK Group (Ichimura, K. et al.). Mitogen-activated protein kinase cascades in plants: a new nomenclature. Trends Plant Sci. 7, 301–308 (2002)
Guo, J. & Chen, J.-G. RACK1 genes regulate plant development with unequal genetic redundancy in Arabidopsis. BMC Plant Biol. 8, 108 (2008)
Li, J.-F. et al. Comprehensive protein-based artificial microRNA screens for effective gene silencing in plants. Plant Cell 25, 1507–1522 (2013)
Witzel, F., Maddison, L. & Blüthgen, N. How scaffolds shape MAPK signaling: what we know and opportunities for systems approaches. Front. Physiol. 3, 475 (2012)
Rahme, L. G. et al. Common virulence factors for bacterial pathogenicity in plants and animals. Science 268, 1899–1902 (1995)
Liberati, N. T. et al. An ordered, nonredundant library of Pseudomonas aeruginosa strain PA14 transposon insertion mutants. Proc. Natl Acad. Sci. USA 103, 2833–2838 (2006)
Motley, S. T. & Lory, S. Functional characterization of a serine/threonine protein kinase of Pseudomonas aeruginosa. Infect. Immun. 67, 5386–5394 (1999)
Parker, J. E., Barber, C. E., Mi-jiao, F. & Daniels, M. J. Interaction of Xanthomonas campestris with Arabidopsis thaliana: characterization of a gene from X. c. pv. raphani that confers avirulence to most A. thaliana accessions. Mol. Plant Microbe Interact. 6, 216–224 (1993)
Djonović, S. et al. Trehalose biosynthesis promotes Pseudomonas aeruginosa pathogenicity in plants. PLoS Pathog. 9, e1003217 (2013)
Prentki, P. & Krisch, H. M. In vitro insertional mutagenesis with a selectable DNA fragment. Gene 29, 303–313 (1984)
Hirsch, A. M. et al. Rhizobium meliloti nodulation genes allow Agrobacterium tumefaciens and Escherichia coli to form pseudonodules on alfalfa. J. Bacteriol. 158, 1133–1143 (1984)
Cheng, Z., Duan, J., Hao, Y., McConkey, B. J. & Glick, B. R. Identification of bacterial proteins mediating the interactions between Pseudomonas putida UW4 and Brassica napus (canola). Mol. Plant Microbe Interact. 22, 686–694 (2009)
Smith, A. W. & Iglewski, B. H. Transformation of Pseudomonas aeruginosa by electroporation. Nucleic Acids Res. 17, 10509 (1989)
Kim, J.-G. et al. Xanthomonas T3S effector XopN suppresses PAMP-triggered immunity and interacts with a tomato atypical receptor-like kinase and TFT1. Plant Cell 21, 1305–1323 (2009)
Curtis, M. D. & Grossniklaus, U. A gateway cloning vector set for high-throughput functional analysis of genes in planta. Plant Physiol. 133, 462–469 (2003)
Lee, L.-Y., Fang, M.-J., Kuang, L.-Y. & Gelvin, S. B. Vectors for multi-color bimolecular fluorescence complementation to investigate protein-protein interactions in living plant cells. Plant Methods 4, 24 (2008)
Zuo, J., Niu, Q. W. & Chua, N. H. Technical advance: an estrogen receptor-based transactivator XVE mediates highly inducible gene expression in transgenic plants. Plant J. 24, 265–273 (2000)
Clough, S. J. & Bent, A. F. Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743 (1998)
Engel, L. S., Hill, J. M., Caballero, A. R., Green, L. C. & O’Callaghan, R. J. Protease IV, a unique extracellular protease and virulence factor from Pseudomonas aeruginosa. J. Biol. Chem. 273, 16792–16797 (1998)
Irizarry, R. A. et al. Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics 4, 249–264 (2003)
Gentleman, R. C. et al. Bioconductor: open software development for computational biology and bioinformatics. Genome Biol. 5, R80 (2004)
Clay, N. K., Adio, A. M., Denoux, C., Jander, G. & Ausubel, F. M. Glucosinolate metabolites required for an Arabidopsis innate immune response. Science 323, 95–101 (2009)
Meyer, D., Lauber, E., Roby, D., Arlat, M. & Kroj, T. Optimization of pathogenicity assays to study the Arabidopsis thaliana–Xanthomonas campestris pv. campestris pathosystem. Mol. Plant Pathol. 6, 327–333 (2005)
Yoo, S. D., Cho, Y. H. & Sheen, J. Arabidopsis mesophyll protoplasts: a versatile cell system for transient gene expression analysis. Nature Protocols 2, 1565–1572 (2007)
Li, J.-F., Bush, J., Xiong, Y., Li, L. & McCormack, M. Large-scale protein-protein interaction analysis in Arabidopsis mesophyll protoplasts by split firefly luciferase complementation. PLoS ONE 6, e27364 (2011)
Li, J.-F., Park, E., von Arnim, A. G. & Nebenfuhr, A. The FAST technique: a simplified Agrobacterium-based transformation method for transient gene expression analysis in seedlings of Arabidopsis and other plant species. Plant Methods 5, 6 (2009)
King, S. R. F. et al. Phytophthora infestans RXLR effector PexRD2 interacts with host MAPKKKε to suppress plant immune signaling. Plant Cell 26, 1345–1359 (2014)
Acknowledgements
We thank G. Tena for generating the mekk1/pMEKK1::MEKK1(K361M) transgenic line, Y. Zhang for the summ1-1 mutant, M. C. Suarez-Rodriguez and P. J. Krysan for discussion, the Arabidopsis Biological Resource Center for tDNA insertion lines, and M. Curtis and U. Grossniklaus for the oestradiol-inducible binary vector. We thank S. Lory for P. aeruginosa PAO ADD1976, and M. B. Mudgett for pVSP61. We thank N. Clay, X. Dong, S. Somerville, and Ausubel laboratory members for reading the manuscript. This work was supported by Natural Sciences and Engineering Research Council of Canada and Banting Postdoctoral Fellowships awarded to Z.C., National Science Foundation grants MCB-0519898 and IOS-0929226 and National Institutes of Health grants R37-GM48707 and P30 DK040561 to F.M.A., and National Science Foundation grant IOS-0618292 and National Institutes of Health grant R01-GM70567 to J.S.
Author information
Authors and Affiliations
Contributions
Z.C., J.-F.L., J.S., and F.M.A. designed experiments, Z.C., J.-F.L., Y.N., X.-C.Z., O.Z.W., Y.X., S.D., Y.M., and J.B. performed experiments, Z.C., J.-F.L., B.J.M., J.S., and F.M.A. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Protease IV-triggered GUS staining in CYP71A12pro:GUS transgenic Arabidopsis seedlings.
a, Activation of CYP71A12pro:GUS by a DEAE fraction of the PA14 secretome (left) and purification of the eliciting activity by DEAE chromatography (right). b, Activation of CYP71A12pro:GUS in 10-day-old seedlings by 100 nM purified PrpL. The experiments in a and b were repeated three times with similar results.
Extended Data Figure 2 Trypsin does not activate MAPK cascade or elicit an oxidative burst in Arabidopsis.
a, Western blot depicting activation of MAPKs by 40 nM flg22, or 40 nM purified PrpL, or trypsin in 10-day-old seedlings. The same molecular mass region of the western blot is shown as in Fig. 1b. b, Chemiluminescence assay showing elicitation of an oxidative burst in 10-day-old seedlings by 20 nM purified PrpL or trypsin. Error bars, s.d.; n = 16 individual seedlings.
Extended Data Figure 3 Transcriptional analysis of purified protease IV.
a, Genome-wide transcriptomic profiles obtained with Affymetrix Arabidopsis ATH1 GeneChips of 10-day-old seedlings treated with 20 nM purified PrpL and comparison with published flg22 and oligogalacturonide responses. A Venn diagram shows the similarity of expression behaviour (|fold change| > 2) in response to the three treatments. b, Defence gene induction levels measured by RT-qPCR in 10-day-old Col-0 seedlings treated with 20 nM purified PrpL or 20 nM flg22 for 1 h (WRKY29, 30, and 33) or 6 h (GST6, ERF1, and CYP71A12). Data represent mean ± s.d.; n = 3 biological replicates, each containing eight seedlings.
Extended Data Figure 4 Protease-IV-triggered responses are dependent on proteolytic activity.
a, Induction of defence-related genes by 20 nM purified PrpL or inactive variants of PrpL measured by RT-qPCR. b, Induction of defence-related genes by 20 nM purified PrpL or 20 nM flg22, or 20 nM TLCK-treated PrpL or 20 nM TLCK-treated flg22 measured by RT-qPCR. c, Induction of defence-related genes by 20 nM PrpL or 20 nM heat-treated PrpL or 20 nM flg22 or 20 nM heat-treated flg22 measured by RT-qPCR. d, Chemiluminescence assay showing elicitation of an oxidative burst by 20 nM purified PrpL, 20 nM inactive variants of PrpL, or 20 nM TLCK-treated PrpL. Data represent mean ± s.d.; n = 3 biological replicates with each experiment contains eight seedlings (a–c) and n = 16 individual seedlings (d).
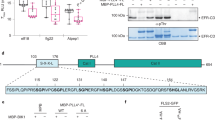
Extended Data Figure 5 Sequence analyses of Xanthomonas argC genes.
a, The protein sequence alignment between P. aeruginosa PA14 PrpL and X. campestris pv. raphani strain 1946 ArgC (Xcr ArgC). b–d, Three independent presumptive null mutations in the Xanthomonas argC gene: an insertion of G, a single nucleotide mutation, and a deletion. The extra G is highlighted in black in b; the single nucleotide substitution is indicated by an arrow in c; and the single base deletion is highlighted in black in d. The resulting premature stop codons are highlighted in red. Sequences were aligned to the argC allele in X. campestris pv. raphani strain 1946 (Xcr-1946), from which the argC gene was cloned. X. campestris pv. campestris strains 8004 (Xcc-8004); X. campestris pv. campestris strains BP109 (Xcc-BP109); X. fuscans subsp. fuscans strain 4834-R (Xf-4834-R); X. campestris pv. vesicatoria (Xcv). e. Activation of CYP71A12pro:GUS in 10-day-old seedlings by culture filtrate from X. campestris strain Xcr-1946, Xcc-8004, or Xcc-BP109, and X. campestris strain 8004 complemented with a functional argC gene (8004/argC) or transformed with empty vector (8004/vector). Detection of HA-ArgC with an anti-HA antibody. The GUS staining was repeated three times with similar results and the representative images shown were selected from at least three images.
Extended Data Figure 6 G proteins are required for protease IV response.
a, Induction of CYP71A12 and GST6 gene expression by 20 nM purified PrpL in 10-day-old wild-type Col-0, gγ single mutants (agg1-1c and agg2-1), or a gγ1γ2 double mutant measured by RT-qPCR. b, Chemiluminescence assay showing elicitation of an oxidative burst by 20 nM purified PrpL or 20 nM flg22 in wild-type Col-0 or G-protein tDNA mutants. Data represent mean ± s.d.; n = 3 biological replicates with each containing eight seedlings (a) and n = 16 individual seedlings (b); P < 0.01, Student’s t-test.
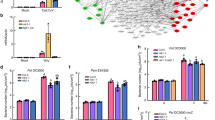
Extended Data Figure 7 Interactions between RACK1 and Gβ or MAPKs.
a, Split-mCherry assay in 4-week-old Agrobacterium-infiltrated N. benthamiana leaves. Images were pseudocoloured for visualization. Scale bar, 100 μm. RACK1A, B, C proteins were fused with the C-terminal half of mCherry and the potential interaction partner proteins were fused with the N-terminal half of mCherry. b, Split-mCherry assay in Arabidopsis protoplasts. RACK1A protein was fused with the C-terminal half of mCherry and the potential interaction partner proteins were fused with the N-terminal half of mCherry. green fluorescent protein (GFP) was included in each experiment to serve as a transfection control. Images were pseudocoloured for visualization. Scale bar, 10 μm. c, Relative interaction intensity between RACK1A and G proteins or MAPKs measured by SFLC. RACK1A protein was fused with the FLucN or FLucC to pair with G proteins or MAPKs fused with the other half of firefly luciferase. Both constructs were co-expressed in protoplasts for 6 h and the complemented luciferase activity was used to relatively quantify protein–protein interactions. UBQ10::GUS was included in each experiment to serve as a transfection normalization control. Data represent mean ± s.d.; n = 3 technical replicate samples. d, Protoplasts were co-transfected with GPA1-HA or AGB1-HA and RACK1B/C-Flag or a control vector. Co-immunoprecipitation was performed with an anti-Flag antibody. Top: the expression of GPA1 or AGB1 protein. Middle: AGB1, but not GPA1, co-immunoprecipitates with RACK1 proteins. Bottom: pulldown of RACK1 proteins by anti-Flag antibody. Protoplasts were treated with 100 nM purified PrpL for 15 min. e, Co-immunoprecipitation between GPA1 or AGB1 and RACK1A was performed in wild-type Col-0 or gαβ mutant Arabidopsis mesophyll protoplasts. Numbers on the left of blots represent marker size in kilodaltons. f, Mass spectrophotometric analysis of endogenous proteins pulled down by Flag-tagged MEKK1(K361M). A peptide conserved in all three RACK1 proteins is shown. The experiments in a and b were repeated three times with similar results.
Extended Data Figure 8 Protease IV-triggered defence responses in wild-type Col-0 and MAPK mutants.
a, Induction of WRKY30 and WRKY33 gene expression by 20 nM purified PrpL in 7-day-old seedlings of wild-type Col-0 and transgenic mpk3,6-es1/2 and mkk4,5-es1/2 plants in the absence or presence of oestradiol. b, Western blot depicting activation of MPK3 and MPK6 by 40 nM purified PrpL in 7-day-old seedlings of wild-type Col-0 and transgenic mkk4,5-es1 plants in the absence or presence of oestradiol. The same molecular mass region of the western blot is shown as in Fig. 1b. c, Induction of WRKY30 and WRKY33 gene expression by 20 nM purified PrpL in 10-day-old wild-type Col-0 and mekk1/pMEKK1::MEKK1(K361M) mutant seedlings. d, Induction of WRKY30 and WRKY33 gene expression by 20 nM purified PrpL in 4-day-old wild-type Col-0 and mekk1 null mutant seedlings. e, Western blot depicting activation of MPK3 and MPK6 by 40 nM purified PrpL in 10-day-old wild-type Col-0 and mekk1/pMEKK1::MEKK1(K361M) mutant seedlings or 4-day-old wild-type Col-0 and mekk1 null mutant seedlings. The same molecular mass region of the western blot is shown as in Fig. 1b. Data represent mean ± s.d.; n = 3 biological replicates with each containing eight seedlings (a, c, d); P < 0.01; P < 0.001, Student’s t-test versus Col-0 controls.
Extended Data Figure 9 RACK1 proteins are required for protease IV response.
a, Western blot depicting activation of MAPKs by 40 nM purified PrpL or 40 nM flg22 in 5-day-old seedlings of wild-type Col-0 and individual rack1::tDNA insertion mutants. The same molecular mass region of the western blot is shown as in Fig. 1b. b, Induction of CYP71A12 by 20 nM purified PrpL or 20 nM flg22 in 5-day-old seedlings of wild-type Col-0 and individual rack1::tDNA insertion mutants. c, RT-qPCR analysis of rack1a, rack1b, and rack1c transcript levels in the 5-day-old Col-0 or amiR–rack1–es1 and amiR–rack1–es2 seedlings. d, RT-qPCR analysis of rack1a, rack1b, and rack1c transcript levels in Arabidopsis protoplasts transfected with amiR–RACK1-4 or artificial microRNA control. e, Western blot depicting activation of MAPKs by 40 nM purified PrpL or 40 nM flg22 in Arabidopsis protoplasts transfected with amiR–RACK1-4 or artificial microRNA control. The same molecular mass region of the western blot is shown as in Fig. 1b. Data represent mean ± s.d.; n = 3 biological replicates (b–d); P < 0.05; P < 0.01, Student’s t-test.
Rights and permissions
About this article
Cite this article
Cheng, Z., Li, JF., Niu, Y. et al. Pathogen-secreted proteases activate a novel plant immune pathway. Nature 521, 213–216 (2015). https://doi.org/10.1038/nature14243
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14243
This article is cited by
-
A critical role of a eubiotic microbiota in gating proper immunocompetence in Arabidopsis
Nature Plants (2023)
-
Mechanisms controlling plant proteases and their substrates
Cell Death & Differentiation (2023)
-
The ColR/S two-component system is a conserved determinant of host association across Pseudomonas species
The ISME Journal (2023)
-
G-protein couples MAPK cascade through maize heterotrimeric Gβ subunit
Plant Cell Reports (2022)
-
CmRCD1 represses flowering by directly interacting with CmBBX8 in summer chrysanthemum
Horticulture Research (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.