Abstract
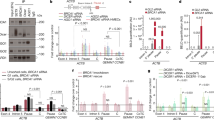
Genome instability is central to ageing, cancer and other diseases. It is not only proteins involved in DNA replication or the DNA damage response (DDR) that are important for maintaining genome integrity: from yeast to higher eukaryotes, mutations in genes involved in pre-mRNA splicing and in the biogenesis and export of messenger ribonucleoprotein (mRNP) also induce DNA damage and genome instability. This instability is frequently mediated by R-loops formed by DNA–RNA hybrids and a displaced single-stranded DNA1. Here we show that the human TREX-2 complex, which is involved in mRNP biogenesis and export, prevents genome instability as determined by the accumulation of γ-H2AX (Ser-139 phosphorylated histone H2AX) and 53BP1 foci and single-cell electrophoresis in cells depleted of the TREX-2 subunits PCID2, GANP and DSS1. We show that the BRCA2 repair factor, which binds to DSS1, also associates with PCID2 in the cell. The use of an enhanced green fluorescent protein-tagged hybrid-binding domain of RNase H1 and the S9.6 antibody did not detect R-loops in TREX-2-depleted cells, but did detect the accumulation of R-loops in BRCA2-depleted cells. The results indicate that R-loops are frequently formed in cells and that BRCA2 is required for their processing. This link between BRCA2 and RNA-mediated genome instability indicates that R-loops may be a chief source of replication stress and cancer-associated instability.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Aguilera, A. & García-Muse, T. R loops: from transcription byproducts to threats to genome stability. Mol. Cell 46, 115–124 (2012)
Huertas, P. & Aguilera, A. Cotranscriptionally formed DNA:RNA hybrids mediate transcription elongation impairment and transcription-associated recombination. Mol. Cell 12, 711–721 (2003)
Tous, C. & Aguilera, A. Impairment of transcription elongation by R-loops in vitro . Biochem. Biophys. Res. Commun. 360, 428–432 (2007)
Kaneko, S., Chu, C., Shatkin, A. & Manley, J. Human capping enzyme promotes formation of transcriptional R loops in vitro . Proc. Natl Acad. Sci. USA 104, 17620–17625 (2007)
Skourti-Stathaki, K., Proudfoot, N. & Gromak, N. Human senataxin resolves RNA/DNA hybrids formed at transcriptional pause sites to promote Xrn2-dependent termination. Mol. Cell 42, 794–805 (2011)
Mischo, H. et al. Yeast Sen1 helicase protects the genome from transcription-associated instability. Mol. Cell 41, 21–32 (2011)
Tuduri, S. et al. Topoisomerase I suppresses genomic instability by preventing interference between replication and transcription. Nature Cell Biol. 11, 1315–1324 (2009)
Wellinger, R., Prado, F. & Aguilera, A. Replication fork progression is impaired by transcription in hyperrecombinant yeast cells lacking a functional THO complex. Mol. Cell. Biol. 26, 3327–3334 (2006)
Gan, W. et al. R-loop-mediated genomic instability is caused by impairment of replication fork progression. Genes Dev. 25, 2041–2056 (2011)
Helmrich, A., Ballarino, M. & Tora, L. Collisions between replication and transcription complexes cause common fragile site instability at the longest human genes. Mol. Cell 44, 966–977 (2011)
Domínguez-Sánchez, M., Barroso, S., Gómez-González, B., Luna, R. & Aguilera, A. Genome instability and transcription elongation impairment in human cells depleted of THO/TREX. PLoS Genet. 7, e1002386 (2011)
Castellano-Pozo, M., García-Muse, T. & Aguilera, A. R-loops cause replication impairment and genome instability during meiosis. EMBO Rep. 13, 923–929 (2012)
Gómez-González, B. et al. Genome-wide function of THO/TREX in active genes prevents R-loop-dependent replication obstacles. EMBO J. 30, 3106–3119 (2011)
Jani, D. et al. Functional and structural characterization of the mammalian TREX-2 complex that links transcription with nuclear messenger RNA export. Nucleic Acids Res. 40, 4562–4573 (2012)
Cabal, G. et al. SAGA interacting factors confine sub-diffusion of transcribed genes to the nuclear envelope. Nature 441, 770–773 (2006)
Umlauf, D. et al. The human TREX-2 complex is stably associated with the nuclear pore basket. J. Cell Sci. 126, 2656–2667 (2013)
Gallardo, M., Luna, R., Erdjument-Bromage, H., Tempst, P. & Aguilera, A. Nab2p and the Thp1p–Sac3p complex functionally interact at the interface between transcription and mRNA metabolism. J. Biol. Chem. 278, 24225–24232 (2003)
Ellisdon, A., Dimitrova, L., Hurt, E. & Stewart, M. Structural basis for the assembly and nucleic acid binding of the TREX-2 transcription-export complex. Nature Struct. Mol. Biol. 19, 328–336 (2012)
Li, J. et al. DSS1 is required for the stability of BRCA2. Oncogene 25, 1186–1194 (2006)
Söderberg, O. et al. Direct observation of individual endogenous protein complexes in situ by proximity ligation. Nature Methods 3, 995–1000 (2006)
El Hage, A., French, S., Beyer, A. & Tollervey, D. Loss of topoisomerase I leads to R-loop-mediated transcriptional blocks during ribosomal RNA synthesis. Genes Dev. 24, 1546–1558 (2010)
Ginno, P., Lott, P., Christensen, H., Korf, I. & Chédin, F. R-loop formation is a distinctive characteristic of unmethylated human CpG island promoters. Mol. Cell 45, 814–825 (2012)
Urtishak, K. et al. Timeless maintains genomic stability and suppresses sister chromatid exchange during unperturbed DNA replication. J. Biol. Chem. 284, 8777–8785 (2009)
Gravells, P. et al. Reduced FANCD2 influences spontaneous SCE and RAD51 foci formation in uveal melanoma and Fanconi anaemia. Oncogene 32, 5338–5346 (2013)
Bermejo, R. et al. The replication checkpoint protects fork stability by releasing transcribed genes from nuclear pores. Cell 146, 233–246 (2011)
Lomonosov, M., Anand, S., Sangrithi, M., Davies, R. & Venkitaraman, A. Stabilization of stalled DNA replication forks by the BRCA2 breast cancer susceptibility protein. Genes Dev. 17, 3017–3022 (2003)
Schlacher, K. et al. Double-strand break repair-independent role for BRCA2 in blocking stalled replication fork degradation by MRE11. Cell 145, 529–542 (2011)
Schlacher, K., Wu, H. & Jasin, M. A distinct replication fork protection pathway connects Fanconi anemia tumor suppressors to RAD51-BRCA1/2. Cancer Cell 22, 106–116 (2012)
Moldovan, G.-L. & D’Andrea, A. How the Fanconi anemia pathway guards the genome. Annu. Rev. Genet. 43, 223–249 (2009)
Yang, H. et al. BRCA2 function in DNA binding and recombination from a BRCA2–DSS1-ssDNA structure. Science 297, 1837–1848 (2002)
ten Asbroek, A., van Groenigen, M., Nooij, M. & Baas, F. The involvement of human ribonucleases H1 and H2 in the variation of response of cells to antisense phosphorothioate oligonucleotides. Eur. J. Biochem. 269, 583–592 (2002)
Bayani, J. & Squire, J. A. Sister chromatid exchange. Curr. Protoc. Cell Biol. 25, 22.7.1–22.7.4 (2005)
Smith, A., Friedman, D., Yu, H., Carnahan, R. & Reynolds, A. ReCLIP (reversible cross-link immuno-precipitation): an efficient method for interrogation of labile protein complexes. PLoS ONE 6, e16206 (2011)
Acknowledgements
We thank J. C. Reyes and A. G. Rondón for comments on the manuscript, and D. Haun for style supervision. Research was funded by grants from the Spanish Ministry of Economy and Competitiveness (Consolider CSD2007-00015 and BFU2010-16372), the Junta de Andalucía (CVI4567) and the European Union (FEDER).
Author information
Authors and Affiliations
Contributions
V.B., S.B., M.G.R., E.T. and E.H.M. performed the experiments. V.B. and A.A. designed the experiments and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Subcellular localization of PCID2.
Immunofluorescence of endogenous PCID2 in HeLa cells. a, Without permeabilization. b, With pre- permeabilization (see Methods).
Extended Data Figure 2 Validation of siRNAs.
a, Relative mRNA quantification. Means and s.e.m. are plotted. b, Western blot analysis of siRNA-treated HeLa cells.
Extended Data Figure 3 Effect of GANP depletion in genomic instability.
a, γ-H2AX and 53BP1 foci. b, Single-cell electrophoresis. c, DNA-combing analysis in GANP-depleted cells. Details as in Fig. 1.
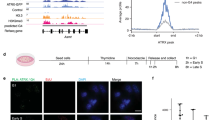
Extended Data Figure 5 HB–GFP interacts with chromatin and chromatin-associated proteins by means of DNA–RNA hybrids.
a, HB–GFP-expressing HEK293 lysate fractionated on Superose 6 size-exclusion columns (17-5172-01; Gelifesciences) and analysed by western blotting. b, HB–GFP co-immunoprecipitated proteins by using the ReCLIP method33. c, HB–GFP ChIP in the ribosomal DNA region in TOP1-depleted HeLa cells (n = 3). Means and s.e.m. are plotted. d, RNH1-dependent release of HB–GFP from chromatin of HeLa cells. e, Scheme and representative plot of FACS assays used to quantify DNA–RNA hybrids. PI, propidium iodide (see Fig. 2f). f, FACS assay to quantify DNA–RNA hybrids in TOP1-depleted cells. Means and s.e.m. are plotted (n = 3).
Extended Data Figure 6 HB–GFP ChIP of the UTRN and ACTB gene regions.
a, b, Normalized siRNA control-relative values for the immunoprecipitated DNA–RNA hybrids plotted relative to the siRNA control in the UTRN (a) and ACTB (b) genes. Means and s.e.m. are shown (n = 3). Amplicon positions used are indicated in the gene diagram above each graph.
Extended Data Figure 7 Accumulation of DNA–RNA hybrids in cells depleted of both PCID2 and BRCA2.
HeLa cells were treated with siBRCA2 in combination with siRNA control or siPCID2, and processed. Means and s.e.m. are plotted (n = 3). Details as in Fig. 3b.
Extended Data Figure 8 Chromosomal aberrations in cells expressing HB–GFP.
a, Metaphase spreads of HeLa cells expressing GFP (control) or HB–GFP. Fragmentation and sister chromatid exchange events are indicated by arrowheads and arrows, respectively. b, Quantification of chromosome breaks in RPE cells expressing HB–GFP. c, Adhesion-independent proliferation assay. Cell proliferation relative to control siRNA-treated RPE cells is shown. Means and s.e.m. are plotted (n = 3). *P ≤ 0.05 (two-tailed Student’s t-test).
Extended Data Figure 9 Model to explain the role of BRCA2 preventing R-loops as a source of genome instability.
a, RNA–DNA hybrids may form both in the interior and at the periphery of the nucleus. mRNP biogenesis factors such as the TREX-2 complex may help recruit or stabilize BRCA2 near transcribed regions, whether or not these are in proximity to the nuclear pore complex. BRCA2 and other related proteins could bind to the branched structure generated by the ssDNA displaced in the R-loop, facilitating the action of enzymes that remove R-loops, such as specific RNases or DNA–RNA helicases. This could occur in non-replicating chromatin. b, In replicating chromatin, BRCA2 and, presumably, other Fanconi anaemia proteins may act directly at putatively stalled RFs in front of an R-loop to impede the collapse or reversal of the replication fork, probably impeding R-loop extension. Subsequently, R-loop removal could be promoted by the passage of the replication fork.
Rights and permissions
About this article
Cite this article
Bhatia, V., Barroso, S., García-Rubio, M. et al. BRCA2 prevents R-loop accumulation and associates with TREX-2 mRNA export factor PCID2. Nature 511, 362–365 (2014). https://doi.org/10.1038/nature13374
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13374
This article is cited by
-
APOBEC3B regulates R-loops and promotes transcription-associated mutagenesis in cancer
Nature Genetics (2023)
-
NKAP acts with HDAC3 to prevent R-loop associated genome instability
Cell Death & Differentiation (2023)
-
miR-4796 enhances the sensitivity of breast cancer cells to ionising radiation by impairing the DNA repair pathway
Breast Cancer (2023)
-
A phase I/II study of preoperative letrozole, everolimus, and carotuximab in stage 2 and 3 hormone receptor-positive and Her2-negative breast cancer
Breast Cancer Research and Treatment (2023)
-
Emerging role for R-loop formation in hepatocellular carcinoma
Genes & Genomics (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.