Abstract
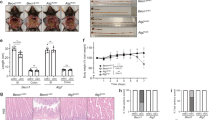
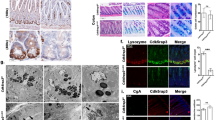
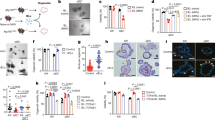
Susceptibility to Crohn’s disease, a complex inflammatory disease involving the small intestine, is controlled by over 30 loci1. One Crohn’s disease risk allele is in ATG16L1, a gene homologous to the essential yeast autophagy gene ATG16 (ref. 2). It is not known how ATG16L1 or autophagy contributes to intestinal biology or Crohn’s disease pathogenesis. To address these questions, we generated and characterized mice that are hypomorphic for ATG16L1 protein expression, and validated conclusions on the basis of studies in these mice by analysing intestinal tissues that we collected from Crohn’s disease patients carrying the Crohn’s disease risk allele of ATG16L1. Here we show that ATG16L1 is a bona fide autophagy protein. Within the ileal epithelium, both ATG16L1 and a second essential autophagy protein ATG5 are selectively important for the biology of the Paneth cell, a specialized epithelial cell that functions in part by secretion of granule contents containing antimicrobial peptides and other proteins that alter the intestinal environment3. ATG16L1- and ATG5-deficient Paneth cells exhibited notable abnormalities in the granule exocytosis pathway. In addition, transcriptional analysis revealed an unexpected gain of function specific to ATG16L1-deficient Paneth cells including increased expression of genes involved in peroxisome proliferator-activated receptor (PPAR) signalling and lipid metabolism, of acute phase reactants and of two adipocytokines, leptin and adiponectin, known to directly influence intestinal injury responses. Importantly, Crohn’s disease patients homozygous for the ATG16L1 Crohn’s disease risk allele displayed Paneth cell granule abnormalities similar to those observed in autophagy-protein-deficient mice and expressed increased levels of leptin protein. Thus, ATG16L1, and probably the process of autophagy, have a role within the intestinal epithelium of mice and Crohn’s disease patients by selective effects on the cell biology and specialized regulatory properties of Paneth cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Barrett, J. C. et al. Genome-wide association defines more than 30 distinct susceptibility loci for Crohn's disease. Nature Genet. 40, 955–962 (2008)
Mizushima, N. et al. Mouse Apg16L, a novel WD-repeat protein, targets to the autophagic isolation membrane with the Apg12–Apg5 conjugate. J. Cell Sci. 116, 1679–1688 (2003)
Ouellette, A. J. Paneth cell α-defensin synthesis and function. Curr. Top. Microbiol. Immunol. 306, 1–25 (2006)
Barbier, M. et al. Overexpression of leptin mRNA in mesenteric adipose tissue in inflammatory bowel diseases. Gastroenterol. Clin. Biol. 27, 987–991 (2003)
Yamamoto, K. et al. Production of adiponectin, an anti-inflammatory protein, in mesenteric adipose tissue in Crohn’s disease. Gut 54, 789–796 (2005)
The Wellcome Trust Case Control Consortium Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature 447, 661–678 (2007)
Xavier, R. J. & Podolsky, D. K. Unravelling the pathogenesis of inflammatory bowel disease. Nature 448, 427–434 (2007)
Rioux, J. D. et al. Genome-wide association study identifies new susceptibility loci for Crohn disease and implicates autophagy in disease pathogenesis. Nature Genet. 39, 596–604 (2007)
Hampe, J. et al. A genome-wide association scan of nonsynonymous SNPs identifies a susceptibility variant for Crohn disease in ATG16L1. Nature Genet. 39, 207–211 (2007)
Levine, B. & Kroemer, G. Autophagy in the Pathogenesis of Disease. Cell 132, 27–42 (2008)
Fujita, N., Itoh, T., Fukuda, M., Noda, T. & Yoshimori, T. The Atg16L complex specifies the site of LC3 lipidation for membrane biogenesis in autophagy. Mol. Biol. Cell 19, 2092–2100 (2008)
Kuma, A., Mizushima, N., Ishihara, N. & Ohsumi, Y. Formation of the approximately 350-kDa Apg12–Apg5.Apg16 multimeric complex, mediated by Apg16 oligomerization, is essential for autophagy in yeast. J. Biol. Chem. 277, 18619–18625 (2002)
Stryke, D. et al. BayGenomics: a resource of insertional mutations in mouse embryonic stem cells. Nucleic Acids Res. 31, 278–281 (2003)
Kuma, A. et al. The role of autophagy during the early neonatal starvation period. Nature 432, 1032–1036 (2004)
Komatsu, M. et al. Homeostatic levels of p62 control cytoplasmic inclusion body formation in autophagy-deficient mice. Cell 131, 1149–1163 (2007)
Hara, T. et al. Suppression of basal autophagy in neural cells causes neurodegenerative disease in mice. Nature 441, 885–889 (2006)
Madison, B. B. et al. Cis elements of the villin gene control expression in restricted domains of the vertical (crypt) and horizontal (duodenum, cecum) axes of the intestine. J. Biol. Chem. 277, 33275–33283 (2002)
Porter, E. M., Bevins, C. L., Ghosh, D. & Ganz, T. The multifaceted Paneth cell. Cell. Mol. Life Sci. 59, 156–170 (2002)
Kobayashi, K. S. et al. Nod2-dependent regulation of innate and adaptive immunity in the intestinal tract. Science 307, 731–734 (2005)
Dvorak, A. M. & Dickersin, G. R. Crohn’s disease: transmission electron microscopic studies I. Barrier function. Possible changes related to alterations of cell coat, mucous coat, epithelial cells, and Paneth cells. Hum. Pathol. 11, 561–571 (1980)
Stappenbeck, T. S., Mills, J. C. & Gordon, J. I. Molecular features of adult mouse small intestinal epithelial progenitors. Proc. Natl Acad. Sci. USA 100, 1004–1009 (2003)
Pua, H. H., Dzhagalov, I., Chuck, M., Mizushima, N. & He, Y. W. A critical role for the autophagy gene Atg5 in T cell survival and proliferation. J. Exp. Med. 204, 25–31 (2007)
Duerr, R. H. et al. A genome-wide association study identifies IL23R as an inflammatory bowel disease gene. Science 314, 1461–1463 (2006)
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005)
Miller, B. C. et al. The autophagy gene ATG5 plays an essential role in B lymphocyte development. Autophagy. 4, 309–314 (2007)
Hara, T. et al. FIP200, a ULK-interacting protein, is required for autophagosome formation in mammalian cells. J. Cell Biol. 181, 497–510 (2008)
Cadwell, K. & Coscoy, L. Ubiquitination on nonlysine residues by a viral E3 ubiquitin ligase. Science 309, 127–130 (2005)
Brown, S. L. et al. Myd88-dependent positioning of Ptgs2-expressing stromal cells maintains colonic epithelial proliferation during injury. J. Clin. Invest. 117, 258–269 (2007)
Pull, S. L., Doherty, J. M., Mills, J. C., Gordon, J. I. & Stappenbeck, T. S. Activated macrophages are an adaptive element of the colonic epithelial progenitor niche necessary for regenerative responses to injury. Proc. Natl Acad. Sci. USA 102, 99–104 (2005)
Acknowledgements
We thank N. Abe for technical assistance with fluorescence microscopy, V. Cavalli for microscope use, J. Eisenberg and A. Ng for help with bioinformatics analysis, A. J. Ouellette for technical advice, and I. Mysorekar and J. Mills for help with antibodies. This research was supported by grant U54 AI057160 Project 6 and the Broad Foundation (K.C., J. Loh and H.W.V.), training grant NIH T32 AR07279 (K.C.), the Lallage Feazel Wall Fellowship DRG-1972-08 (K.C.) from the Damon Runyon Cancer Research Foundation; the Pew Foundation (J. Liu, S.L.B., H.M., W.K. and T.S.S.); the Washington University Digestive Diseases Research Core Center P30 DK52574, Barnes Jewish Foundation, Johnson and Johnson Translational Seed Award, and the Crohn’s Colitis Foundation of America (E.L., S.H. and C.S.); grants-in-aid for Scientific Research from the Ministry of Education, Culture, Sports, Science and Technology of Japan, and the Toray Science Foundation (C.K. and N.M.); NIH R01 AI062832 (J.A.C.); and NIH AI062773 and DK43351 (R.J.X.).
Author Contributions The original hypothesis of the article was formulated by K.C., H.W.V. and T.S.S. Autophagy experiments were performed by K.C. and C.K. Mouse lines were established by K.C., J. Loh and B.P.S. Mouse analysis experiments were performed by K.C., J. Liu, S.L.B., H.M., W.K., T.S.S. and J. Lennerz. Histological examination was performed by T.S.S, E.M.B. and J. Lennerz. L. monocytogenes experiments were performed by J.C. and K.C. Microarray analysis was performed by R.J.X. and T.S.S. Human tissue collection, genotyping and preparation were performed by E.L., S.H. and C.S. N.M. provided advice, the antibody to ATG16L1, and experiments by C.K. were performed in N.M.’s laboratory. The manuscript was written by K.C., H.W.V. and T.S.S., and all authors commented on the manuscript, data and conclusions before submission.
Author information
Authors and Affiliations
Corresponding authors
Supplementary information
Supplementary Figures
This file contains Supplementary Figures 1-14 with Legends (PDF 9724 kb)
Supplementary Table
Supplementary Table 1. Microarray analysis of RNA from crypt-base epithelial cells (XLS 58 kb)
Rights and permissions
About this article
Cite this article
Cadwell, K., Liu, J., Brown, S. et al. A key role for autophagy and the autophagy gene Atg16l1 in mouse and human intestinal Paneth cells. Nature 456, 259–263 (2008). https://doi.org/10.1038/nature07416
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature07416
This article is cited by
-
Transcriptome-wide association studies associated with Crohn’s disease: challenges and perspectives
Cell & Bioscience (2024)
-
Loss of Mptx2 alters bacteria composition and intestinal homeostasis potentially by impairing autophagy
Communications Biology (2024)
-
Inner mitochondrial membrane protein Prohibitin 1 mediates Nix-induced, Parkin-independent mitophagy
Scientific Reports (2023)
-
Scribble deficiency mediates colon inflammation by inhibiting autophagy-dependent oxidative stress elimination
Scientific Reports (2023)
-
Bioengineering translational models of lymphoid tissues
Nature Reviews Bioengineering (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.