Abstract
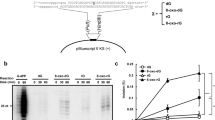
DNA lesions can often block DNA replication, so cells possess specialized low-fidelity, and often error-prone, DNA polymerases that can bypass such lesions and promote replication of damaged DNA1. The Saccharomyces cerevisiae RAD30 and human hRAD30A encode Polη, which bypasses a cis–syn thymine–thymine dimer efficiently and accurately2,3,4,5,6,7. Here we show that a related human gene, hRAD30B8, encodes the DNA polymerase Polι, which misincorporates deoxynucleotides at a high rate. To bypass damage, Polι specifically incorporates deoxynucleotides opposite highly distorting or non-instructional DNA lesions. This action is combined with that of DNA polymerase Polζ, which is essential for damage-induced mutagenesis, to complete the lesion bypass. Polζ is very inefficient in inserting deoxynucleotides opposite DNA lesions, but readily extends from such deoxynucleotides once they have been inserted. Thus, in a new model for mutagenic bypass of DNA lesions in eukaryotes, the two DNA polymerases act sequentially: Polι incorporates deoxynucleotides opposite DNA lesions, and Polζ functions as a mispair extender.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Johnson, R. E., Washington, M. T., Prakash, S. & Prakash, L. Bridging the gap: a family of novel DNA polymerases that replicate faulty DNA. Proc. Natl Acad. Sci. USA 96, 12224 –12226 (1999).
Johnson, R. E., Prakash, S. & Prakash, L. Efficient bypass of a thymine-thymine dimer by yeast DNA polymerase, Polη. Science 283, 1001 –1004 (1999).
Johnson, R. E., Kondratick, C. M., Prakash, S. & Prakash, L. hRAD30 mutations in the variant form of xeroderm pigmentosum. Science 285, 263–265 ( 1999).
Masutani, C. et al. The XPV (xeroderma pigmentosum variant) gene encodes human DNA polymerase η. Nature 399, 700–704 (1999).
Washington, M. T., Johnson, R. E., Prakash, S. & Prakash, L. Accuracy of thymine-thymine dimer bypass by Saccharomyces cerevisiae DNA polymerase η. Proc. Natl Acad. Sci. USA 97, 3094–3099 (2000).
Johnson, R. E., Washington, M. T., Prakash, S. & Prakash, L. Fidelity of human DNA polymerase η. J. Biol. Chem. 275, 7447–7450 (2000).
Washington, M. T., Johnson, R. E., Prakash, S. & Prakash, L. Fidelity and processivity of Saccharomyces cerevisiae DNA polymerase η. J. Biol. Chem. 274, 36835– 36838 (1999).
McDonald, J. P. et al. Novel human and mouse homologs of Saccharomyces cerevisiae DNA polymerase η. Genomics 60, 20–30 (1999).
Goodman, M. F., Creighton, S., Bloom, L. B. & Petruska, J. Biochemical basis of DNA replication fidelity. Crit. Rev. Biochem. Mol. Biol. 28, 83–126 ( 1993).
Creighton, S., Bloom, L. B. & Goodman, M. F. Gel fidelity assay measuring nucleotide misinsertion, exonucleolytic proofreading, and lesion bypass efficiencies. Methods Enzymol. 262, 232–256 ( 1995).
Nelson, J. R., Lawrence, C. W. & Hinkle, D. C. Thymine-thymine dimer bypass by yeast DNA polymerase ζ. Science 272, 1646–1649 (1996).
Gibbs, P. E. M., McGregor, W. G., Maher, V. M., Nisson, P. & Lawrence, C. W. A human homolog of the Saccharomyces cerevisiae REV3 gene, which encodes the catalytic subunit of DNA polyerase ζ. Proc. Natl Acad. Sci. USA 95, 6876– 6880 (1998).
Thomas, D. C. et al. Fidelity of mammalian DNA replication and replicative DNA polymerases. Biochem. 30, 11751– 11759 (1991).
Mendelman, L. V., Petruska, J. & Goodman, M. F. Base mispair extension kinetics. Comparison of DNA polymerase α and reverse transcriptase. J. Biol. Chem. 265, 2338–2346 (1990).
Gibbs, P. E. M., Kilbey, B. J., Banerjee, S. K. & Lawrence, C. W. The frequency and accuracy of replication past a thymine-thymine cyclobutane dimer are very different in Saccharomyces cerevisiae and Escherichia coli. J. Bacteriol. 175, 2607– 2612 (1993).
Gibbs, P. E. M., Borden, A. & Lawrence, C. W. The T–T pyrimidine (6-4) pyrimidinone UV photoproduct is much less mutagenic in yeast than in Escherichia coli. Nucleic Acids Res. 23, 1919–1922 (1995).
Ciarrocchi, G. & Pedrini, A. M. Determination of pyrimidine dimer unwinding angle by measurement of DNA electrophoretic mobility. J. Mol. Biol. 155, 177– 183 (1982).
Kim, J.-K., Patel, D. & Choi, B.-S. Contrasting structural impacts induced by cis–syn cyclobutane dimer and (6-4) adduct in DNA duplex decamers: implication in mutagenesis and repair activity. Photochem. Photobiol. 62, 44–50 (1995).
Kim, J.-K. & Choi, B.-S. The solution structure of DNA duplex-decamer containing the (6-4) photoproduct of thymidyl(3′→5′)thymidine by NMR and relaxation matrix refinement. Eur. J. Biochem. 228, 849–854 (1995).
Wong, I., Patel, S. S. & Johnson, K. A. An induced-fit kinetic mechanism for DNA replication fidelity: direct measurement by single-turnover kinetics. Biochemistry 30, 526–537 ( 1991).
Huang, M.-M., Arnheim, N. & Goodman, M. F. Extension of base mispairs by Taq DNA polymerase: implications for single nucleotide discrimination in PCR. Nucleic Acids Res. 20, 4567–4573 (1992).
Acknowledgements
We thank D. Hinkle for plasmids pGST-Rev3 and pRev7 and R. Hodge for providing the T–T dimer- and (6-4) photoproduct-containing DNAs, the construction of which was supported by a National Institute of Environmental Health Science (NIEHS) Centre Grant. This work was supported by a grant from the NIH.
Author information
Additional information
Sealy Centre for Molecular Science, University of Texas Medical Branch at Galveston, 6.104 Medical Research Building, 11th and Mechanic Streets, Galveston, Texas 77555-1061, USA
Supplementary information
Rights and permissions
About this article
Cite this article
Johnson, R., Washington, M., Haracska, L. et al. Eukaryotic polymerases ι and ζ act sequentially to bypass DNA lesions. Nature 406, 1015–1019 (2000). https://doi.org/10.1038/35023030
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/35023030
This article is cited by
-
Mechanism of nucleotide discrimination by the translesion synthesis polymerase Rev1
Nature Communications (2022)
-
Genetic and physical interactions between Polη and Rev1 in response to UV-induced DNA damage in mammalian cells
Scientific Reports (2021)
-
Alcohol-derived DNA crosslinks are repaired by two distinct mechanisms
Nature (2020)
-
Structure of DNA polymerase ζ: capturing the getaway driver
Nature Structural & Molecular Biology (2020)
-
Distinct requirements for budding yeast Rev1 and Polη in translesion DNA synthesis across different types of DNA damage
Current Genetics (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.