Key Points
-
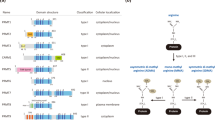
The royal superfamily domains, which include Tudor, chromo, MBT, PWWP and agenet domains, read protein methylation. The Tudor protein family includes a group of proteins that are specialized to recognize arginine methylation, which is an ability that has not been found in other royal superfamily domains.
-
The Tudor family proteins are classified into methylarginine-binding and methyllysine-binding groups. Most methylarginine-binding Tudor proteins are associated with RNA metabolism and most methyllysine-binding Tudor proteins are implicated in chromatin biology.
-
The Tudor domain core contains a conserved β-barrel structure, with an aromatic cage for methyl-ligand recognition. Crystal structures of ligand-bound extended Tudor domains (eTuds) have provided insights into the structural mechanism of methylarginine reading. Tudor domain–methylarginine binding interactions have been implicated in biological processes such as mRNA splicing, small RNA pathways and transciptional regulation.
-
Arginine methylation is an evolutionarily conserved mark on PIWI proteins and is read by the eTuds of the germline Tudor proteins. Methylation-dependent Tudor–PIWI interactions are required for the proper function of the PIWI-interacting RNA (piRNA) pathway.
-
In the animal germ line, genetic studies have revealed that Tudor proteins function as adaptors to facilitate the assembly of nuage and ensure undisrupted gametogenesis.
Abstract
Proteins can be modified by post-translational modifications such as phosphorylation, methylation, acetylation and ubiquitylation, creating binding sites for specific protein domains. Methylation has pivotal roles in the formation of complexes that are involved in cellular regulation, including in the generation of small RNAs. Arginine methylation was discovered half a century ago, but the ability of methylarginine sites to serve as binding motifs for members of the Tudor protein family, and the functional significance of the protein–protein interactions that are mediated by Tudor domains, has only recently been appreciated. Tudor proteins are now known to be present in PIWI complexes, where they are thought to interact with methylated PIWI proteins and regulate the PIWI-interacting RNA (piRNA) pathway in the germ line.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Seet, B. T., Dikic, I., Zhou, M. M. & Pawson, T. Reading protein modifications with interaction domains. Nature Rev. Mol. Cell Biol. 7, 473–483 (2006).
Boswell, R. E. & Mahowald, A. P. Tudor, a gene required for assembly of the germ plasm in Drosophila melanogaster. Cell 43, 97–104 (1985).
Ponting, C. P. Tudor domains in proteins that interact with RNA. Trends Biochem. Sci. 22, 51–52 (1997).
Callebaut, I. & Mornon, J. P. The human EBNA-2 coactivator p100: multidomain organization and relationship to the staphylococcal nuclease fold and to the Tudor protein involved in Drosophila melanogaster development. Biochem. J. 321, 125–132 (1997). References 3 and 4 were the first papers to define the Tudor domain.
Buhler, D., Raker, V., Luhrmann, R. & Fischer, U. Essential role for the Tudor domain of SMN in spliceosomal U snRNP assembly: implications for spinal muscular atrophy. Hum. Mol. Genet. 8, 2351–2357 (1999).
Selenko, P. et al. SMN Tudor domain structure and its interaction with the Sm proteins. Nature Struct. Biol. 8, 27–31 (2001). This study reports the first Tudor domain structure, describing the signature β-barrel fold and aromatic cage.
Friesen, W. J., Massenet, S., Paushkin, S., Wyce, A. & Dreyfuss, G. SMN, the product of the spinal muscular atrophy gene, binds preferentially to dimethylarginine-containing protein targets. Mol. Cell 7, 1111–1117 (2001). A seminal work showing that the Tudor domain-containing protein SMN can bind methylated-arginine targets.
Sprangers, R., Groves, M. R., Sinning, I. & Sattler, M. High-resolution X-ray and NMR structures of the SMN Tudor domain: conformational variation in the binding site for symmetrically dimethylated arginine residues. J. Mol. Biol. 327, 507–520 (2003).
Maurer-Stroh, S. et al. The Tudor domain 'Royal Family': Tudor, plant Agenet, Chromo, PWWP and MBT domains. Trends Biochem. Sci. 28, 69–74 (2003). This paper defines the royal superfamily domains.
Taverna, S. D., Li, H., Ruthenburg, A. J., Allis, C. D. & Patel, D. J. How chromatin-binding modules interpret histone modifications: lessons from professional pocket pickers. Nature Struct. Mol. Biol. 14, 1025–1040 (2007).
Adams-Cioaba, M. A. & Min, J. Structure and function of histone methylation binding proteins. Biochem. Cell Biol. 87, 93–105 (2009).
Yap, K. L. & Zhou, M. M. Keeping it in the family: diverse histone recognition by conserved structural folds. Crit. Rev. Biochem. Mol. Biol. 45, 488–505 (2010).
Kim, J. et al. Tudor, MBT and chromo domains gauge the degree of lysine methylation. EMBO Rep. 7, 397–403 (2006).
Ruthenburg, A. J., Li, H., Patel, D. J. & Allis, C. D. Multivalent engagement of chromatin modifications by linked binding modules. Nature Rev. Mol. Cell Biol. 8, 983–994 (2007).
Jin, J. et al. Eukaryotic protein domains as functional units of cellular evolution. Sci. Signal. 2, ra76 (2009).
Siomi, M. C., Mannen, T. & Siomi, H. How does the royal family of Tudor rule the PIWI-interacting RNA pathway? Genes Dev. 24, 636–646 (2010). An excellent review on Tudor–PIWI interactions.
Rogne, M. et al. The KH-Tudor domain of A-kinase anchoring protein 149 mediates RNA-dependent self-association. Biochemistry 45, 14980–14989 (2006).
Botuyan, M. V. et al. Structural basis for the methylation state-specific recognition of histone H4-K20 by 53BP1 and Crb2 in DNA repair. Cell 127, 1361–1373 (2006).
Huang, Y., Fang, J., Bedford, M. T., Zhang, Y. & Xu, R. M. Recognition of histone H3 lysine-4 methylation by the double Tudor domain of JMJD2A. Science 312, 748–751 (2006).
Liu, H. et al. Structural basis for methylarginine-dependent recognition of Aubergine by Tudor. Genes Dev. 24, 1876–1881 (2010).
Liu, K. et al. Structural basis for recognition of arginine methylated Piwi proteins by the extended Tudor domain. Proc. Natl Acad. Sci. USA 107, 18398–18403 (2010). References 20 and 21 describe the first crystal structures of Tudor domains in complex with methylated arginine-containing peptides.
Shaw, N. et al. The multifunctional human p100 protein 'hooks' methylated ligands. Nature Struct. Mol. Biol. 14, 779–784 (2007).
Li, C. L., Yang, W. Z., Chen, Y. P. & Yuan, H. S. Structural and functional insights into human Tudor-SN, a key component linking RNA interference and editing. Nucleic Acids Res. 36, 3579–3589 (2008).
Friberg, A., Corsini, L., Mourao, A. & Sattler, M. Structure and ligand binding of the extended Tudor domain of D. melanogaster Tudor-SN. J. Mol. Biol. 387, 921–934 (2009).
Pahlich, S., Zakaryan, R. P. & Gehring, H. Protein arginine methylation: cellular functions and methods of analysis. Biochim. Biophys. Acta 1764, 1890–1903 (2006).
Brahms, H., Meheus, L., de Brabandere, V., Fischer, U. & Luhrmann, R. Symmetrical dimethylation of arginine residues in spliceosomal Sm protein B/B′ and the Sm-like protein LSm4, and their interaction with the SMN protein. RNA 7, 1531–1542 (2001).
Cote, J. & Richard, S. Tudor domains bind symmetrical dimethylated arginines. J. Biol. Chem. 280, 28476–28483 (2005). This paper reveals the general concept of Tudor domains as methylarginine readers.
Friesen, W. J. & Dreyfuss, G. Specific sequences of the Sm and Sm-like (Lsm) proteins mediate their interaction with the spinal muscular atrophy disease gene product (SMN). J. Biol. Chem. 275, 26370–26375 (2000).
Brahms, H. et al. The C-terminal RG dipeptide repeats of the spliceosomal Sm proteins D1 and D3 contain symmetrical dimethylarginines, which form a major B-cell epitope for anti-Sm autoantibodies. J. Biol. Chem. 275, 17122–17129 (2000).
Talbot, K., Miguel-Aliaga, I., Mohaghegh, P., Ponting, C. P. & Davies, K. E. Characterization of a gene encoding survival motor neuron (SMN)-related protein, a constituent of the spliceosome complex. Hum. Mol. Genet. 7, 2149–2156 (1998).
Meister, G. et al. Methylation of Sm proteins by a complex containing PRMT5 and the putative U snRNP assembly factor pICln. Curr. Biol. 11, 1990–1994 (2001).
Rappsilber, J., Ajuh, P., Lamond, A. I. & Mann, M. SPF30 is an essential human splicing factor required for assembly of the U4/U5/U6 tri-small nuclear ribonucleoprotein into the spliceosome. J. Biol. Chem. 276, 31142–31150 (2001).
Anne, J. Arginine methylation of SmB is required for Drosophila germ cell development. Development 137, 2819–2828 (2010).
Cheng, D., Cote, J., Shaaban, S. & Bedford, M. T. The arginine methyltransferase CARM1 regulates the coupling of transcription and mRNA processing. Mol. Cell 25, 71–83 (2007).
Yang, Y. et al. TDRD3 is an effector molecule for arginine-methylated histone marks. Mol. Cell 40, 1016–1023 (2010).
Sims, R. J., et al. The C-terminal domain of RNA polymerase II is modified by site-specific methylation. Science 332, 99–103 (2011).
Linder, B. et al. Tdrd3 is a novel stress granule-associated protein interacting with the Fragile-X syndrome protein FMRP. Hum. Mol. Genet. 17, 3236–3246 (2008).
Goulet, I., Boisvenue, S., Mokas, S., Mazroui, R. & Cote, J. TDRD3, a novel Tudor domain-containing protein, localizes to cytoplasmic stress granules. Hum. Mol. Genet. 17, 3055–3074 (2008).
Chen, C. et al. Mouse Piwi interactome identifies binding mechanism of Tdrkh Tudor domain to arginine methylated Miwi. Proc. Natl Acad. Sci. USA 106, 20336–20341 (2009).
Kirino, Y. et al. Arginine methylation of Piwi proteins catalysed by dPRMT5 is required for Ago3 and Aub stability. Nature Cell Biol. 11, 652–658 (2009). Provides the first evidence that arginine methylation is an evolutionarily conserved modification on PIWI proteins.
Kojima, K. et al. Associations between PIWI proteins and TDRD1/MTR-1 are critical for integrated subcellular localization in murine male germ cells. Genes Cells 14, 1155–1165 (2009).
Nishida, K. M. et al. Functional involvement of Tudor and dPRMT5 in the piRNA processing pathway in Drosophila germlines. EMBO J. 28, 3820–3831 (2009).
Reuter, M. et al. Loss of the Mili-interacting Tudor domain-containing protein-1 activates transposons and alters the Mili-associated small RNA profile. Nature Struct. Mol. Biol. 16, 639–646 (2009).
Vagin, V. V. et al. Proteomic analysis of murine Piwi proteins reveals a role for arginine methylation in specifying interaction with Tudor family members. Genes Dev. 23, 1749–1762 (2009). References 39 and 44 are two proteomic studies that highlight the fact that germline Tudor domain proteins are physiological binding partners of PIWI family proteins.
Wang, J., Saxe, J. P., Tanaka, T., Chuma, S. & Lin, H. Mili interacts with Tudor domain-containing protein 1 in regulating spermatogenesis. Curr. Biol. 19, 640–644 (2009).
Kirino, Y. et al. Arginine methylation of Aubergine mediates Tudor binding and germ plasm localization. RNA 16, 70–78 (2010).
Gonsalvez, G. B., Rajendra, T. K., Tian, L. & Matera, A. G. The Sm-protein methyltransferase, dart5, is essential for germ-cell specification and maintenance. Curr. Biol. 16, 1077–1089 (2006).
Anne, J., Ollo, R., Ephrussi, A. & Mechler, B. M. Arginine methyltransferase Capsuleen is essential for methylation of spliceosomal Sm proteins and germ cell formation in Drosophila. Development 134, 137–146 (2007).
Kirino, Y. et al. Arginine methylation of Vasa protein is conserved across phyla. J. Biol. Chem. 285, 8148–8154 (2010).
Arkov, A. L. & Ramos, A. Building RNA-protein granules: insight from the germline. Trends Cell Biol. 20, 482–490 (2010).
Aravin, A. A. et al. Cytoplasmic compartmentalization of the fetal piRNA pathway in mice. PLoS Genet. 5, e1000764 (2009).
Thomson, T. & Lasko, P. Tudor and its domains: germ cell formation from a Tudor perspective. Cell Res. 15, 281–291 (2005).
Arkov, A. L., Wang, J. Y., Ramos, A. & Lehmann, R. The role of Tudor domains in germline development and polar granule architecture. Development 133, 4053–4062 (2006).
Aravin, A. A. et al. Double-stranded RNA-mediated silencing of genomic tandem repeats and transposable elements in the D. melanogaster germline. Curr. Biol. 11, 1017–1027 (2001).
Vagin, V. V. et al. The RNA interference proteins and vasa locus are involved in the silencing of retrotransposons in the female germline of Drosophila melanogaster. RNA Biol. 1, 54–58 (2004).
Lim, A. K. & Kai, T. Unique germ-line organelle, nuage, functions to repress selfish genetic elements in Drosophila melanogaster. Proc. Natl Acad. Sci. USA 104, 6714–6719 (2007).
Malone, C. D. et al. Specialized piRNA pathways act in germline and somatic tissues of the Drosophila ovary. Cell 137, 522–535 (2009).
Patil, V. S. & Kai, T. Repression of retroelements in Drosophila germline via piRNA pathway by the Tudor domain protein Tejas. Curr. Biol. 20, 724–730 (2010).
Liu, L., Qi, H., Wang, J. & Lin, H. PAPI, a novel TUDOR-domain protein, complexes with AGO3, ME31B and TRAL in the nuage to silence transposition. Development 138, 1863–1873 (2011).
Szakmary, A., Reedy, M., Qi, H. & Lin, H. The Yb protein defines a novel organelle and regulates male germline stem cell self-renewal in Drosophila melanogaster. J. Cell Biol. 185, 613–627 (2009).
Olivieri, D., Sykora, M. M., Sachidanandam, R., Mechtler, K. & Brennecke, J. An in vivo RNAi assay identifies major genetic and cellular requirements for primary piRNA biogenesis in Drosophila. EMBO J. 29, 3301–3317 (2010).
Saito, K. et al. Roles for the Yb body components Armitage and Yb in primary piRNA biogenesis in Drosophila. Genes Dev. 24, 2493–2498 (2010).
Qi, H. et al. The Yb body, a major site for Piwi-associated RNA biogenesis and a gateway for Piwi expression and transport to the nucleus in somatic cells. J. Biol. Chem. 286, 3789–3797 (2011).
Chuma, S. et al. Tdrd1/Mtr-1, a Tudor-related gene, is essential for male germ-cell differentiation and nuage/germinal granule formation in mice. Proc. Natl Acad. Sci. USA 103, 15894–15899 (2006).
Kuramochi-Miyagawa, S. et al. Mili, a mammalian member of piwi family gene, is essential for spermatogenesis. Development 131, 839–849 (2004).
Carmell, M. A. et al. MIWI2 is essential for spermatogenesis and repression of transposons in the mouse male germline. Dev. Cell 12, 503–514 (2007).
Pan, J. et al. RNF17, a component of the mammalian germ cell nuage, is essential for spermiogenesis. Development 132, 4029–4039 (2005).
Vasileva, A., Tiedau, D., Firooznia, A., Muller-Reichert, T. & Jessberger, R. Tdrd6 is required for spermiogenesis, chromatoid body architecture, and regulation of miRNA expression. Curr. Biol. 19, 630–639 (2009).
Lachke, S. A. et al. Mutations in the RNA granule component TDRD7 cause cataract and glaucoma. Science 331, 1571–1576 (2011).
Tanaka, T. et al. Tudor domain containing 7 (Tdrd7) is essential for dynamic ribonucleoprotein (RNP) remodeling of chromatoid bodies during spermatogenesis. Proc. Natl Acad. Sci. USA 108, 10579–10584 (2011).
Deng, W. & Lin, H. miwi, a murine homolog of piwi, encodes a cytoplasmic protein essential for spermatogenesis. Dev. Cell 2, 819–830 (2002).
Hosokawa, M. et al. Tudor-related proteins TDRD1/MTR-1, TDRD6 and TDRD7/TRAP: domain composition, intracellular localization, and function in male germ cells in mice. Dev. Biol. 301, 38–52 (2007).
Yabuta, Y. et al. TDRD5 is required for retrotransposon silencing, chromatoid body assembly, and spermiogenesis in mice. J. Cell Biol. 192, 781–795 (2011).
Shoji, M. et al. The TDRD9–MIWI2 complex is essential for piRNA-mediated retrotransposon silencing in the mouse male germline. Dev. Cell 17, 775–787 (2009).
Tee, W. W. et al. Prmt5 is essential for early mouse development and acts in the cytoplasm to maintain ES cell pluripotency. Genes Dev. 24, 2772–2777 (2010).
Pawson, T. & Scott, J. D. Signaling through scaffold, anchoring, and adaptor proteins. Science 278, 2075–2080 (1997).
Pawson, T. Dynamic control of signaling by modular adaptor proteins. Curr. Opin. Cell Biol. 19, 112–116 (2007).
Matthews, A. G. et al. RAG2 PHD finger couples histone H3 lysine 4 trimethylation with V(D)J recombination. Nature 450, 1106–1110 (2007).
Zhao, Q. et al. PRMT5-mediated methylation of histone H4R3 recruits DNMT3A, coupling histone and DNA methylation in gene silencing. Nature Struct. Mol. Biol. 16, 304–311 (2009).
Gary, J. D. & Clarke, S. RNA and protein interactions modulated by protein arginine methylation. Prog. Nucleic Acid Res. Mol. Biol. 61, 65–131 (1998).
Bedford, M. T. & Richard, S. Arginine methylation an emerging regulator of protein function. Mol. Cell 18, 263–272 (2005).
Bedford, M. T. & Clarke, S. G. Protein arginine methylation in mammals: who, what, and why. Mol. Cell 33, 1–13 (2009).
Chang, B., Chen, Y., Zhao, Y. & Bruick, R. K. JMJD6 is a histone arginine demethylase. Science 318, 444–447 (2007).
Webby, C. J. et al. Jmjd6 catalyses lysyl-hydroxylation of U2AF65, a protein associated with RNA splicing. Science 325, 90–93 (2009).
Kim, V. N., Han, J. & Siomi, M. C. Biogenesis of small RNAs in animals. Nature Rev. Mol. Cell Biol. 10, 126–139 (2009).
Siomi, M. C., Sato, K., Pezic, D. & Aravin, A. A. PIWI-interacting small RNAs: the vanguard of genome defence. Nature Rev. Mol. Cell Biol. 12, 246–258 (2011).
Hutvagner, G. & Simard, M. J. Argonaute proteins: key players in RNA silencing. Nature Rev. Mol. Cell Biol. 9, 22–32 (2008).
Aravin, A. et al. A novel class of small RNAs bind to MILI protein in mouse testes. Nature 442, 203–207 (2006).
Grivna, S. T., Beyret, E., Wang, Z. & Lin, H. A novel class of small RNAs in mouse spermatogenic cells. Genes Dev. 20, 1709–1714 (2006).
Girard, A., Sachidanandam, R., Hannon, G. J. & Carmell, M. A. A germline-specific class of small RNAs binds mammalian Piwi proteins. Nature 442, 199–202 (2006).
Lau, N. C. et al. Characterization of the piRNA complex from rat testes. Science 313, 363–367 (2006).
Watanabe, T. et al. Identification and characterization of two novel classes of small RNAs in the mouse germline: retrotransposon-derived siRNAs in oocytes and germline small RNAs in testes. Genes Dev. 20, 1732–1743 (2006).
Aravin, A. A. & Hannon, G. J. Small RNA silencing pathways in germ and stem cells. Cold Spring Harb. Symp. Quant. Biol. 73, 283–290 (2008).
Aravin, A. A., Hannon, G. J. & Brennecke, J. The Piwi-piRNA pathway provides an adaptive defense in the transposon arms race. Science 318, 761–764 (2007).
Brennecke, J. et al. Discrete small RNA-generating loci as master regulators of transposon activity in Drosophila. Cell 128, 1089–1103 (2007).
Gunawardane, L. S. et al. A slicer-mediated mechanism for repeat-associated siRNA 5′ end formation in Drosophila. Science 315, 1587–1590 (2007).
Vagin, V. V., Hannon, G. J. & Aravin, A. A. Arginine methylation as a molecular signature of the Piwi small RNA pathway. Cell Cycle 8, 4003–4004 (2009).
Lau, N. C. Small RNAs in the animal gonad: guarding genomes and guiding development. Int. J. Biochem. Cell Biol. 42, 1334–1347 (2010).
Jacobs, S. A. & Khorasanizadeh, S. Structure of HP1 chromodomain bound to a lysine 9-methylated histone H3 tail. Science 295, 2080–2083 (2002).
Min, J. et al. L3MBTL1 recognition of mono- and dimethylated histones. Nature Struct. Mol. Biol. 14, 1229–1230 (2007).
Vezzoli, A. et al. Molecular basis of histone H3K36me3 recognition by the PWWP domain of Brpf1. Nature Struct. Mol. Biol. 17, 617–619 (2010).
Ramos, A. et al. The structure of the N-terminal domain of the fragile X mental retardation protein: a platform for protein–protein interaction. Structure 14, 21–31 (2006).
Acknowledgements
We dedicate this paper to Maggie Pawson, in memory of her many contributions to the world of signal transduction. We thank J. Park for assistance with the figures and B. Liu, G. Gish and J. Min for comments on the manuscript. C.C. and J.J. are recipients of fellowships from the Canadian Institutes for Health Research (CIHR). Work in the laboratory of T.P. is supported by grants from the CIHR, the Canadian Cancer Society, Genome Canada through the Ontario Genomics Institute and the Ontario Research Fund.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
41580_2011_BFnrm3185_MOESM1_ESM.pdf
Supplementary information S1 (timeline) | Historical timeline of Tudor domain recognition of arginine methylation. (PDF 256 kb)
41580_2011_BFnrm3185_MOESM2_ESM.pdf
Supplementary information S2 (figure) | Domain architectures of the Tudor domain protein family of human, Drosophila and fission yeast. (PDF 289 kb)
41580_2011_BFnrm3185_MOESM4_ESM.pdf
Supplementary information S4 (table) | Function of the mammalian TDRD group of Tudor proteins in the germline. (PDF 241 kb)
Glossary
- Chromo
-
A royal superfamily domain of ∼50 amino acids that is best known for binding to methylated lysine on histone tails.
- MBT
-
A royal superfamily domain that binds to methylated lysine.
- PWWP
-
A royal superfamily domain that contains a PWWP signature motif. Some PWWP domains bind methylated lysine.
- Plant agenet
-
A Tudor-like domain that was discovered in plants and named after the Plantagenet dynasty of English monarchs. This royal superfamily domain is also present in animal proteins such as fragile X mental retardation protein (FMRP) and its autosomal paralogues fragile X mental retardation syndrome-related protein 1 (FXR1) and FXR2.
- Transposon
-
A mobile DNA element that can translocate in the genome. Transposons are considered as parasitic DNA elements of the host genome and can be classified based on the mechanism of transposition as retrotransposons (copy and paste) and DNA transposons (cut and paste).
- Plant homeodomain
-
(PHD). PHD domains are conserved zinc-finger domains that are found in many nuclear proteins involved in chromatin remodelling and bind methylated lysines on histone tails.
- WD40
-
A protein motif of ∼40 amino acids that usually ends with a WD dipeptide. Repeated WD40 motifs fold into a circular β-propeller structure that mediates protein–protein interactions.
- K homology
-
(KH). A protein fold of ∼70 amino acids that was first identified in heterologous nuclear ribonucleoprotein K (HNRNPK). KH domains bind RNA or single-stranded DNA and are present in a wide array of nucleic acid-binding proteins.
- Staphylococcal nuclease-like
-
(SN-like). The SN-like domain is implicated in single-stranded nucleic acid-binding and is present as four intact repeats in SN-like domain containing protein 1(SND1).
- Cation–π interactions
-
Noncovalent electrostatic interactions between an electron-rich π system (for example, in an aromatic ring) and a cation.
- Small nuclear ribonucleoprotein
-
(snRNP). RNA–protein complexes that are components of the spliceosome, which is the precursor mRNA splicing apparatus found in eukaryotic nuclei.
- Polar granules
-
Maternally produced cytoplasmic granules localized at the posterior end of a Drosophila melanogaster early embryo and oocyte. These granules are required for germ cell formation.
- Nuage
-
A perinuclear, electron-dense, cloud-like structure first identified in Drosophila melanogaster ovarian nurse cells. It is used as a general term for germ granules, which are cytoplasmic, non-membranous, ribonucleoprotein organelles that are found in animal germ cells.
- Long interspersed nuclear elements
-
(LINEs). A family of autonomous non-long terminal repeat (non-LTR) retrotransposons that mobilize by retrotransposition. They are one of the most abundant groups of retrotransposons in the human genome.
- Pi-body
-
Pi-bodies are intermitochondrial cement-like, cytoplasmic germ granules that are found in mammalian fetal spermatogonia. The proteins that are found in the pi-body include several PIWI-interacting RNA pathway proteins such as TDRD1 (Tudor domain-containing protein 1), MILI, Vasa, GASZ (germ cell-specific ankyrin, SAM and basic leucine zipper domain-containing protein) and MOV10L1 (moloney leukaemia virus 10-like protein 1).
- PiP-body
-
Processing body (P-body)-like bodies are cytoplasmic germ granules in fetal spermatogonia that are distinct from pi-bodies. PiP-bodies contain PIWI-interacting RNA pathway components such as MIWI2, Tudor-domain containing protein 9 (TDRD9), Maelstrom (MAEL) and Vasa, and also processing-body markers such as GW182, mRNA-decapping enzyme 1A (DCP1A), 5–3′ exoribonuclease 1 (XRN1) and DEAD box protein 6 (DDX6).
- Chromatoid body
-
A prominent perinuclear electron-dense germ granule in male postmeiotic round spermatids. It is enriched with RNA and protein components of the PIWI-interacting RNA pathway as well as those of the microRNA pathway.
- Lotus domain
-
(Also known as an OST-HTH domain). A conserved protein fold of ∼90 amino acids that is present in several germline- specific proteins, including Linkman, Oskar and Tudor domain-containing protein 5 (TDRD5) and TDRD7. The lotus domain is speculated to bind RNA.
- RNA recognition motif
-
(RRM). A protein fold of ∼90 amino acids with putative RNA-binding properties that is present in many RNA-binding proteins.
- Intermitochondrial cement
-
An electron-dense cementing material that is associated with mitochondria in mammalian spermatocytes and spermatogonia.
Rights and permissions
About this article
Cite this article
Chen, C., Nott, T., Jin, J. et al. Deciphering arginine methylation: Tudor tells the tale. Nat Rev Mol Cell Biol 12, 629–642 (2011). https://doi.org/10.1038/nrm3185
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrm3185
This article is cited by
-
Keep quiet: the HUSH complex in transcriptional silencing and disease
Nature Structural & Molecular Biology (2024)
-
Development and validation of an RBP gene signature for prognosis prediction in colorectal cancer based on WGCNA
Hereditas (2023)
-
Carcinogenesis promotion in oral squamous cell carcinoma: KDM4A complex-mediated gene transcriptional suppression by LEF1
Cell Death & Disease (2023)
-
The Fragile X Protein Family in Amyotrophic Lateral Sclerosis
Molecular Neurobiology (2023)
-
Evolutionary dynamics and conserved function of the Tudor domain-containing (TDRD) proteins in teleost fish
Marine Life Science & Technology (2022)