Abstract
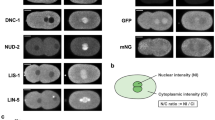
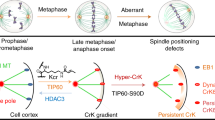
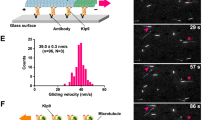
Despite being essential for spatial cell division control, the mechanisms governing spindle positioning remain incompletely understood. In the Caenorhabditis elegans one-cell stage embryo, the spindle becomes asymmetrically positioned during anaphase through the action of as-yet unidentified cortical force generators that pull on astral microtubules and that depend on two Gα proteins and associated proteins1,2. We performed spindle-severing experiments following temporally restricted gene inactivation and drug exposure, and established that microtubule dynamics and dynein are both required for generating efficient pulling forces. We found that the Gα-associated proteins GPR-1/2 and LIN-5 interact in vivo with LIS-1, a component of the dynein complex. Moreover, we discovered that the LIN-5, GPR-1/2 and the Gα proteins promote the presence of the dynein complex at the cell cortex. Our findings suggest a mechanism by which the Gα proteins enable GPR-1/2 and LIN-5 recruitment to the cortex, thus ensuring the presence of cortical dynein. Together with microtubule dynamics, this allows pulling forces to be exerted and proper cell division to be achieved.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Colombo, K. et al. Translation of polarity cues into asymmetric spindle positioning in Caenorhabditis elegans embryos. Science 300, 1957–1961 (2003).
Grill, S. W., Howard, J., Schaffer, E., Stelzer, E. H. & Hyman, A. A. The distribution of active force generators controls mitotic spindle position. Science 301, 518–521 (2003).
Grill, S. W., Gönczy, P., Stelzer, E. H. & Hyman, A. A. Polarity controls forces governing asymmetric spindle positioning in the Caenorhabditis elegans embryo. Nature 409, 630–633 (2001).
Gotta, M. & Ahringer, J. Distinct roles for Galpha and Gbetagamma in regulating spindle position and orientation in Caenorhabditis elegans embryos. Nature Cell Biol. 3, 297–300 (2001).
Gotta, M., Dong, Y., Peterson, Y. K., Lanier, S. M. & Ahringer, J. Asymmetrically distributed C. elegans homologs of AGS3/PINS control spindle position in the early embryo. Curr. Biol. 13, 1029–1037 (2003).
Srinivasan, D. G., Fisk, R. M., Xu, H. & van den Heuvel, S. A complex of LIN-5 and GPR proteins regulates G protein signaling and spindle function in C. elegans. Genes Dev. 17, 1225–1239 (2003).
Lorson, M. A., Horvitz, H. R. & van den Heuvel, S. LIN-5 is a novel component of the spindle apparatus required for chromosome segregation and cleavage plane specification in Caenorhabditis elegans. J. Cell Biol. 148, 73–86 (2000).
Afshar, K., Willard, F. S., Colombo, K., Siderovski, D. P. & Gönczy, P. Cortical localization of the Galpha protein GPA-16 requires RIC-8 function during C. elegans asymmetric cell division. Development 132, 4449–ß4459 (2005).
Bellaiche, Y. & Gotta, M. Heterotrimeric G proteins and regulation of size asymmetry during cell division. Curr. Opin. Cell Biol. 17, 658–663 (2005).
Du, Q. & Macara, I. G. Mammalian Pins is a conformational switch that links NuMA to heterotrimeric G proteins. Cell 119, 503–516 (2004).
Izumi, Y., Ohta, N., Hisata, K., Raabe, T. & Matsuzaki, F. Drosophila Pins-binding protein Mud regulates spindle-polarity coupling and centrosome organization. Nature Cell Biol. 8, 586–593 (2006).
Siller, K. H., Cabernard, C. & Doe, C. Q. The NuMA-related Mud protein binds Pins and regulates spindle orientation in Drosophila neuroblasts. Nature Cell Biol. 8, 594–600 (2006).
Bowman, S. K., Neumuller, R. A., Novatchkova, M., Du, Q. & Knoblich, J. A. The Drosophila NuMA Homolog Mud regulates spindle orientation in asymmetric cell division. Dev. Cell 10, 731–742 (2006).
Kozlowski, C., Srayko, M. & Nedelec, F. Cortical microtubule contacts position the spindle in C. elegans embryos. Cell 129, 499–510 (2007).
Wright, A. J. & Hunter, C. P. Mutations in a beta-tubulin disrupt spindle orientation and microtubule dynamics in the early Caenorhabditis elegans embryo. Mol. Biol. Cell 14, 4512–4525 (2003).
Hyman, A. A. & White, J. G. Determination of cell division axes in the early embryogenesis of Caenorhabditis elegans. J. Cell Biol. 105, 2123–2135 (1987).
Severson, A. F. & Bowerman, B. Myosin and the PAR proteins polarize microfilament-dependent forces that shape and position mitotic spindles in Caenorhabditis elegans. J. Cell Biol. 161, 21–26 (2003).
Schmidt, D. J., Rose, D. J., Saxton, W. M. & Strome, S. Functional analysis of cytoplasmic dynein heavy chain in Caenorhabditis elegans with fast-acting temperature-sensitive mutations. Mol. Biol. Cell 16, 1200–1212 (2005).
Gönczy, P., Pichler, S., Kirkham, M. & Hyman, A. A. Cytoplasmic dynein is required for distinct aspects of MTOC positioning, including centrosome separation, in the one cell stage Caenorhabditis elegans embryo. J. Cell Biol. 147, 135–150 (1999).
Hamill, D. R., Severson, A. F., Carter, J. C. & Bowerman, B. Centrosome maturation and mitotic spindle assembly in C. elegans require SPD-5, a protein with multiple coiled-coil domains. Dev. Cell 3, 673–684 (2002).
Encalada, S. E., Willis, J., Lyczak, R. & Bowerman, B. A spindle checkpoint functions during mitosis in the early Caenorhabditis elegans embryo. Mol. Biol. Cell 16, 1056–1070 (2005).
Merdes, A., Ramyar, K., Vechio, J. D. & Cleveland, D. W. A complex of NuMA and cytoplasmic dynein is essential for mitotic spindle assembly. Cell 87, 447–458 (1996).
Cockell, M. M., Baumer, K. & Gönczy, P. lis-1 is required for dynein-dependent cell division processes in C. elegans embryos. J. Cell Sci. 117, 4571–4582 (2004).
Vallee, R. B., Tai, C. & Faulkner, N. E. LIS1: cellular function of a disease-causing gene. Trends Cell Biol. 11, 155–160 (2001).
Carminati, J. L. & Stearns, T. Microtubules orient the mitotic spindle in yeast through dynein-dependent interactions with the cell cortex. J. Cell Biol. 138, 629–641 (1997).
Gupta, M. L. Jr., Carvalho, P., Roof, D. M. & Pellman, D. Plus end-specific depolymerase activity of Kip3, a kinesin-8 protein, explains its role in positioning the yeast mitotic spindle. Nature Cell Biol. 8, 913–923 (2006).
Cottingham, F. R. & Hoyt, M. A. Mitotic spindle positioning in Saccharomyces cerevisiae is accomplished by antagonistically acting microtubule motor proteins. J. Cell Biol. 138, 1041–1053 (1997).
Fink, G., Schuchardt, I., Colombelli, J., Stelzer, E. & Steinberg, G. Dynein-mediated pulling forces drive rapid mitotic spindle elongation in Ustilago maydis. EMBO J. 25, 4897–4908 (2006).
Pecreaux, J. et al. Spindle oscillations during asymmetric cell division require a threshold number of active cortical force generators. Curr. Biol. 16, 2111–2122 (2006).
Mallik, R., Carter, B. C., Lex, S. A., King, S. J. & Gross, S. P. Cytoplasmic dynein functions as a gear in response to load. Nature 427, 649–652 (2004).
Daniels, B. R., Masi, B. C. & Wirtz, D. Probing single-cell micromechanics in vivo: the microrheology of C. elegans developing embryos. Biophys. J. 90, 4712–4719 (2006).
Strome, S. et al. Spindle dynamics and the role of gamma-tubulin in early Caenorhabditis elegans embryos. Mol. Biol. Cell 12, 1751–1764 (2001).
Schlaitz, A. L. et al. The C. elegans RSA complex localizes protein phosphatase 2A to centrosomes and regulates mitotic spindle assembly. Cell 128, 115–127 (2007).
Afshar, K. et al. RIC-8 is required for GPR-1/2-dependent Galpha function during asymmetric division of C. elegans embryos. Cell 119, 219–230 (2004).
Gönczy, P. et al. Dissection of cell division processes in the one cell stage Caenorhabditis elegans embryo by mutational analysis. J. Cell Biol. 144, 927–946 (1999).
Acknowledgements
We are grateful to Daniel Constam, Virginie Hachet and Kalyani Thyagarajan for critical reading of the manuscript, Virginie Hachet for affinity purifying GPR-1/2 antibodies, John Blanc and Hugues Baumgartner for building the temperature-shift device, Claude Bonnard and Natsuko Imaizumi for help with image processing. Mutant and transgenic strains were kindly provided by Bruce Bowerman, Craig Hunter, Anthony Hyman and Sander van den Heuvel. Supported by grant 3100A0-102087 from the Swiss National Science Foundation.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
Supplementary Figures S1,S2, S3, S4, S5, S6, S7, S8 and S9, Supplementary Movie Legends and Supplementary Tables 1 and 2 (PDF 1944 kb)
Rights and permissions
About this article
Cite this article
Nguyen-Ngoc, T., Afshar, K. & Gönczy, P. Coupling of cortical dynein and Gα proteins mediates spindle positioning in Caenorhabditis elegans. Nat Cell Biol 9, 1294–1302 (2007). https://doi.org/10.1038/ncb1649
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb1649
This article is cited by
-
Laser ablation and fluid flows reveal the mechanism behind spindle and centrosome positioning
Nature Physics (2024)
-
AMBRA1 phosphorylation by CDK1 and PLK1 regulates mitotic spindle orientation
Cellular and Molecular Life Sciences (2023)
-
A force-balance model for centrosome positioning and spindle elongation during interphase and anaphase B
Indian Journal of Physics (2022)
-
Spindle positioning and its impact on vertebrate tissue architecture and cell fate
Nature Reviews Molecular Cell Biology (2021)
-
Cytoskeletal control of early mammalian development
Nature Reviews Molecular Cell Biology (2021)