Abstract
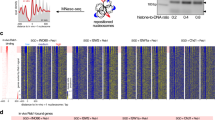
The ATP-dependent chromatin assembly and remodelling factor (ACF) functions to generate regularly spaced nucleosomes, which are required for heritable gene silencing. The mechanism by which ACF mobilizes nucleosomes remains poorly understood. Here we report a single-molecule FRET study that monitors the remodelling of individual nucleosomes by ACF in real time, revealing previously unknown remodelling intermediates and dynamics. In the presence of ACF and ATP, the nucleosomes exhibit gradual translocation along DNA interrupted by well-defined kinetic pauses that occurred after approximately seven or three to four base pairs of translocation. The binding of ACF, translocation of DNA and exiting of translocation pauses are all ATP-dependent, revealing three distinct functional roles of ATP during remodelling. At equilibrium, a continuously bound ACF complex can move the nucleosome back-and-forth many times before dissociation, indicating that ACF is a highly processive and bidirectional nucleosome translocase.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Becker, P. B. & Horz, W. ATP-dependent nucleosome remodeling. Annu. Rev. Biochem. 71, 247–273 (2002)
Narlikar, G. J., Fan, H. Y. & Kingston, R. E. Cooperation between complexes that regulate chromatin structure and transcription. Cell 108, 475–487 (2002)
Flaus, A. & Owen-Hughes, T. Mechanisms for ATP-dependent chromatin remodelling: farewell to the tuna-can octamer? Curr. Opin. Genet. Dev. 14, 165–173 (2004)
Smith, C. L. & Peterson, C. L. ATP-dependent chromatin remodeling. Curr. Top. Dev. Biol. 65, 115–148 (2004)
Clapier, C. R. & Cairns, B. R. The biology of chromatin remodeling complexes. Annu. Rev. Biochem. 78, 273–304 (2009)
Saha, A., Wittmeyer, J. & Cairns, B. R. Chromatin remodeling by RSC involves ATP-dependent DNA translocation. Genes Dev. 16, 2120–2134 (2002)
Whitehouse, I., Stockdale, C., Flaus, A., Szczelkun, M. D. & Owen-Hughes, T. Evidence for DNA translocation by the ISWI chromatin-remodeling enzyme. Mol. Cell. Biol. 23, 1935–1945 (2003)
Tsukiyama, T., Palmer, J., Landel, C. C., Shiloach, J. & Wu, C. Characterization of the Imitation Switch subfamily of ATP-dependent chromatin-remodeling factors in Saccharomyces cerevisiae . Genes Dev. 13, 686–697 (1999)
Hamiche, A., Sandaltzopoulos, R., Gdula, D. A. & Wu, C. ATP-dependent histone octamer sliding mediated by the chromatin remodeling complex NURF. Cell 97, 833–842 (1999)
Längst, G., Bonte, E. J., Corona, D. F. & Becker, P. B. Nucleosome movement by CHRAC and ISWI without disruption or trans-displacement of the histone octamer. Cell 97, 843–852 (1999)
Kassabov, S. R., Henry, N. M., Zofall, M., Tsukiyama, T. & Bartholomew, B. High-resolution mapping of changes in histone-DNA contacts of nucleosomes remodeled by ISW2. Mol. Cell. Biol. 22, 7524–7534 (2002)
Zhang, Y. et al. DNA translocation and loop formation mechanism of chromatin remodeling by SWI/SNF and RSC. Mol. Cell 24, 559–568 (2006)
Lia, G. et al. Direct observation of DNA distortion by the RSC complex. Mol. Cell 21, 417–425 (2006)
Shundrovsky, A., Smith, C. L., Lis, J. T., Peterson, C. L. & Wang, M. D. Probing SWI/SNF remodeling of the nucleosome by unzipping single DNA molecules. Nature Struct. Mol. Biol. 13, 549–554 (2006)
Stryer, L. & Haugland, R. P. Energy transfer: a spectroscopic ruler. Proc. Natl Acad. Sci. USA 58, 719–726 (1967)
Ha, T. et al. Probing the interaction between two single molecules: fluorescence resonance energy transfer between a single donor and a single acceptor. Proc. Natl Acad. Sci. USA 93, 6264–6268 (1996)
Zhuang, X. et al. A single molecule study of RNA catalysis and folding. Science 288, 2048–2051 (2000)
Ito, T., Bulger, M., Pazin, M. J., Kobayashi, R. & Kadonaga, J. T. ACF, an ISWI-containing and ATP-utilizing chromatin assembly and remodeling factor. Cell 90, 145–155 (1997)
Ito, T. et al. ACF consists of two subunits, Acf1 and ISWI, that function cooperatively in the ATP-dependent catalysis of chromatin assembly. Genes Dev. 13, 1529–1539 (1999)
Bochar, D. A. et al. A family of chromatin remodeling factors related to Williams syndrome transcription factor. Proc. Natl Acad. Sci. USA 97, 1038–1043 (2000)
LeRoy, G., Loyola, A., Lane, W. S. & Reinberg, D. Purification and characterization of a human factor that assembles chromatin. J. Biol. Chem. 275, 14787–14790 (2000)
Poot, R. A. et al. HuCHRAC, a human ISWI chromatin remodelling complex contains hACF1 and two novel histone-fold proteins. EMBO J. 19, 3377–3387 (2000)
Yang, J. G., Madrid, T. S., Sevastopoulos, E. & Narlikar, G. J. The chromatin-remodeling enzyme ACF is an ATP-dependent DNA length sensor that regulates nucleosome spacing. Nature Struct. Mol. Biol. 13, 1078–1083 (2006)
Lowary, P. T. & Widom, J. New DNA sequence rules for high affinity binding to histone octamer and sequence-directed nucleosome positioning. J. Mol. Biol. 276, 19–42 (1998)
Abbondanzieri, E. A. et al. Dynamic binding orientations direct activity of HIV reverse transcriptase. Nature 453, 184–189 (2008)
He, X., Fan, H. Y., Narlikar, G. J. & Kingston, R. E. Human ACF1 alters the remodeling strategy of SNF2h. J. Biol. Chem. 281, 28636–28647 (2006)
Stockdale, C., Flaus, A., Ferreira, H. & Owen-Hughes, T. Analysis of nucleosome repositioning by yeast ISWI and Chd1 chromatin remodeling complexes. J. Biol. Chem. 281, 16279–16288 (2006)
Schwanbeck, R., Xiao, H. & Wu, C. Spatial contacts and nucleosome step movements induced by the NURF chromatin remodeling complex. J. Biol. Chem. 279, 39933–39941 (2004)
Zofall, M., Persinger, J., Kassabov, S. R. & Bartholomew, B. Chromatin remodeling by ISW2 and SWI/SNF requires DNA translocation inside the nucleosome. Nature Struct. Mol. Biol. 13, 339–346 (2006)
Dang, W. & Bartholomew, B. Domain architecture of the catalytic subunit in the ISW2-nucleosome complex. Mol. Cell. Biol. 27, 8306–8317 (2007)
Li, G., Levitus, M., Bustamante, C. & Widom, J. Rapid spontaneous accessibility of nucleosomal DNA. Nature Struct. Mol. Biol. 12, 46–53 (2004)
Partensky, P. D. & Narlikar, G. J. Chromatin remodelers act globally, sequence positions nucleosomes locally. J. Mol. Biol. 391, 12–25 (2009)
Luger, K., Mader, A. W., Richmond, R. K., Sargent, D. F. & Richmond, T. J. Crystal structure of the nucleosome core particle at 2.8 Å resolution. Nature 389, 251–260 (1997)
Fyodorov, D. V. & Kadonaga, J. T. Dynamics of ATP-dependent chromatin assembly by ACF. Nature 418, 896–900 (2002)
Gangaraju, V. K., Prasad, P., Srour, A., Kagalwala, M. N. & Bartholomew, B. Conformational changes associated with template commitment in ATP-dependent chromatin remodeling by ISW2. Mol. Cell 35, 58–69 (2009)
Raki, L. R. The chromatin remodeller ACF acts as a dimeric motor to space nucleosomes. Nature 10.1038/nature08621 (this issue)
Strohner, R. et al. A ‘loop recapture’ mechanism for ACF-dependent nucleosome remodeling. Nature Struct. Mol. Biol. 12, 683–690 (2005)
Fitzgerald, D. J. et al. Reaction cycle of the yeast Isw2 chromatin remodeling complex. EMBO J. 23, 3836–3843 (2004)
Cairns, B. R. Chromatin remodeling: insights and intrigue from single-molecule studies. Nature Struct. Mol. Biol. 14, 989–996 (2007)
Luger, K., Rechsteiner, T. J. & Richmond, T. J. Preparation of nucleosome core particle from recombinant histones. Methods Enzymol. 304, 3–19 (1999)
Aalfs, J. D., Narlikar, G. J. & Kingston, R. E. Functional differences between the human ATP-dependent nucleosome remodeling proteins BRG1 and SNF2H. J. Biol. Chem. 276, 34270–34278 (2001)
Rasnik, I., McKinney, S. A. & Ha, T. Nonblinking and long-lasting single-molecule fluorescence imaging. Nature Methods 3, 891–893 (2006)
Acknowledgements
We thank J. Widom for providing the plasmid containing the 601 positioning sequence and R. E. Kingston for the plasmids containing the SNF2h and Acf1 genes. We also thank L. Racki and E. Abbondanzieri for helpful discussions, and W. Huang and B. Harada for help with some experiments. This work is supported in part by Howard Hughes Medical Institute (to X.Z.) and the National Institutes of Health (GM073767) and the Beckman Foundation (to G.J.N). X.Z. is a Howard Hughes Medical Institute investigator. M.D.S. was a NIH Ruth L. Kirschstein NSRA Fellow, G.J.N is a Leukemia and Lymphoma Society Scholar.
Author Contributions T.R.B. performed the experiments and analysis with help from M.D.S.; J.G.Y. made the enzymes and histone proteins. T.R.B., G.J.N. and X.Z. designed the experiments. X.Z. oversaw the project.
Author information
Authors and Affiliations
Corresponding authors
Supplementary information
Supplementary Figures
This file contains Supplementary Figures 1-12 with Legends. (PDF 6014 kb)
Rights and permissions
About this article
Cite this article
Blosser, T., Yang, J., Stone, M. et al. Dynamics of nucleosome remodelling by individual ACF complexes. Nature 462, 1022–1027 (2009). https://doi.org/10.1038/nature08627
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature08627
This article is cited by
-
Changes in histone lysine acetylation, but not DNA methylation during facultative hibernation in Syrian hamster liver
Epigenetics Communications (2024)
-
Functionalized graphene-oxide grids enable high-resolution cryo-EM structures of the SNF2h-nucleosome complex without crosslinking
Nature Communications (2024)
-
Nucleosome density shapes kilobase-scale regulation by a mammalian chromatin remodeler
Nature Structural & Molecular Biology (2023)
-
Single-Molecule Characterization of Cy3.5 -Cy5.5 Dye Pair for FRET Studies of Nucleic Acids and Nucleosomes
Journal of Fluorescence (2023)
-
H2A.Z deposition by SWR1C involves multiple ATP-dependent steps
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.